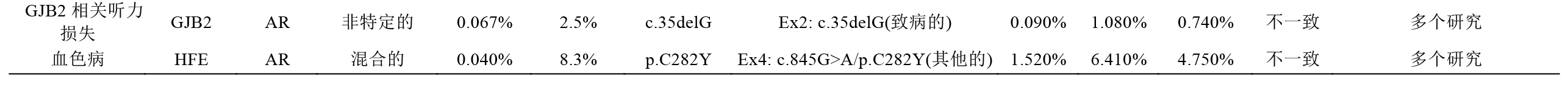目录
遗传变异分类标准与指南
免责声明
These ACMG Standards and Guidelines were developed primarily as an educational resource for clinical laboratory geneticists to help them provide quality clinical laboratory services. Adherence to these standards and guidelines is voluntary and does not necessarily assure a successful medical outcome. These Standards and Guidelines should not be considered inclusive of all proper procedures and tests or exclusive of other procedures and tests that are reasonably directed to obtaining the same results. In determining the propriety of any specific procedure or test, the clinical laboratory geneticist should apply his or her own professional judgment to the specific circumstances presented by the individual patient or specimen. Clinical laboratory geneticists are encouraged to document in the patient’s record the rationale for the use of a particular procedure or test, whether or not it is in conformance with these Standards and Guidelines. They also are advised to take notice of the date any particular guideline was adopted and to consider other relevant medical and scientific information that becomes available after that date. It also would be prudent to consider whether intellectual property interests may restrict the performance of certain tests and other procedures.
ACMG制定的标准与指南作为教育资源旨在帮助临床遗传学家提供优质的临床检验服务。遵循该标准和指南属于自愿行为并且不一定能够确保一个成功的医疗结局。该标准和指南并不囊括所有合适的流程和检测,也不排斥其他可以获得相同结果的流程和检测。临床实验室遗传学家应该利用自己的专业知识,依据病人或样本的具体情况来判断某一具体的流程或检测的合理性。我们鼓励临床实验室遗传学家记录对病人使用的某一具体流程或检测的原理,不管这个原理与这些标准与指南是否符合。同时建议临床实验室遗传学家关注指南的采用时间,应考虑到此后更新的一些相关医疗和科学信息。还需谨慎考虑到知识产权可能会限制某些检测或流程的使用。
摘要
The American College of Medical Genetics and Genomics (ACMG) previously developed guidance for the interpretation of sequence variants.1 In the past decade, sequencing technology has evolved rapidly with the advent of high-throughput next-generation sequencing. By adopting and leveraging next-generation sequencing, clinical laboratories are now performing an ever-increasing catalogue of genetic testing spanning genotyping, single genes, gene panels, exomes, genomes, transcriptomes, and epigenetic assays for genetic disorders. By virtue of increased complexity, this shift in genetic testing has been accompanied by new challenges in sequence interpretation. In this context the ACMG convened a workgroup in 2013 comprising representatives from the ACMG, the Association for Molecular Pathology (AMP), and the College of American Pathologists to revisit and revise the standards and guidelines for the interpretation of sequence variants. The group consisted of clinical laboratory directors and clinicians. This report represents expert opinion of the workgroup with input from ACMG, AMP, and College of American Pathologists stakeholders. These recommendations primarily apply to the breadth of genetic tests used in clinical laboratories, including genotyping, single genes, panels, exomes, and genomes. This report recommends the use of specific standard terminology—“pathogenic,” “likely pathogenic,” “uncertain significance,” “likely benign,” and “benign”—to describe variants identified in genes that cause Mendelian disorders. Moreover, this recommendation describes a process for classifying variants into these five categories based on criteria using typical types of variant evidence (e.g., population data, computational data, functional data, segregation data). Because of the increased complexity of analysis and interpretation of clinical genetic testing described in this report, the ACMG strongly recommends that clinical molecular genetic testing should be performed in a Clinical Laboratory Improvement Amendments–approved laboratory, with results interpreted by a board-certified clinical molecular geneticist or molecular genetic pathologist or the equivalent.
美国医学遗传学与基因组学学会(The American College of Medical Genetics and Genomics,ACMG)曾制定过序列变异解读指南。在过去的十年中,随着新一代高通量测序的出现,测序技术有了快速发展。利用新一代测序技术,临床实验室检测遗传性疾病的产品种类不断增加,包括基因分型、单基因、基因包、外显子组、基因组、转录组和表观遗传学检测。随着技术的复杂性日益增加,基因检测在序列解读方面不断面临着新的挑战。因此ACMG在2013年成立了一个工作组来重新审视和修订序列变异解读的标准和指南,工作组包括ACMG、分子病理协会(the Association for Molecular Pathology,AMP)和美国病理学家协会(the College of American Pathologists,CAP)的代表。该工作组由临床实验室主任和临床医生组成。本报告代表了工作组中来自ACMG,AMP和CAP的专家意见。本报告提出的建议可应用于临床实验室的各种基因检测方法,包括基因分型、单基因、基因包、外显子组和基因组。本报告建议使用特定标准术语来描述孟德尔疾病相关的基因变异——“致病的”、“可能致病的”、“意义不明确的”、“可能良性的”和“良性的”。此外,本报告描述了基于典型的数据类型(如人群数据,计算数据,功能数据,共分离数据)对变异进行五级分类的标准过程。由于临床基因检测分析和解读中不断增加的复杂性,ACMG强烈建议临床分子基因检测应在符合临床实验室改进修正案(CLIA)认证的实验室中进行,其检测结果应由通过职业认证的临床分子遗传学家或分子遗传病理学家或相同职能的专业人员解读。
Key Words 关键词
ACMG laboratory guideline; clinical genetic testing; interpretation; reporting; sequence variant terminology; variant reporting
ACMG实验室指南;临床遗传检测;解读;报告;序列变异术语;变异报告
1.引言
Clinical molecular laboratories are increasingly detecting novel sequence variants in the course of testing patient specimens for a rapidly increasing number of genes associated with genetic disorders. While some phenotypes are associated with a single gene, many are associated with multiple genes. Our understanding of the clinical significance of any given sequence variant falls along a gradient, ranging from those in which the variant is almost certainly pathogenic for a disorder to those that are almost certainly benign. While the previous American College of Medical Genetics and Genomics (ACMG) recommendations provided interpretative categories of sequence variants and an algorithm for interpretation, the recommendations did not provide defined terms or detailed variant classification guidance.1 This report describes updated standards and guidelines for the classification of sequence variants using criteria informed by expert opinion and empirical data.
随着遗传病患者样本中所检测基因数目的快速增加,临床分子实验室检测到越来越多的新的序列变异。某些表型仅与单个基因相关,而多数表型与多个基因相关。对某个给定序列变异的临床意义进行分级解读,从某个变异几乎可以肯定是某种疾病的致病性变异到几乎可以肯定是良性变异。虽然ACMG之前的建议提供了序列变异的解读分类及解读的算法,但并没有提供明确的术语或详细的变异分类指导。本研究依据专家意见及经验数据,阐述了最新的序列变异分类标准和指南。
2.方法
In 2013 a workgroup consisting of ACMG, Association for Molecular Pathology (AMP), and College of American Pathologists members, representing clinical laboratory directors and clinicians, was formed with the goal of developing a recommendation for the use of standard terminology for classifying sequence variants using available evidence weighted according to a system developed through expert opinion, workgroup consensus, and community input. To assess the views of the clinical laboratory community, surveys were sent to over 100 sequencing laboratories in the United States and Canada that were listed in GeneTests.org, requesting input on terminology preferences and evaluation of evidence for classifying variants. Laboratory testing experience included rare disease as well as pharmacogenomics and somatic cancer testing. The first survey, aimed at assessing terminology preferences, was sent in February 2013, and the results were presented in an open forum at the 2013 ACMG annual meeting including over 75 attendees. Survey respondents represented more than 45 laboratories in North America. The outcome of the survey and open forum indicated that (i) a five-tier terminology system using the terms “pathogenic,” “likely pathogenic,” “uncertain significance,” “likely benign,” and “benign” was preferred and already in use by a majority of laboratories, and (ii) the first effort of the workgroup should focus on Mendelian and mitochondrial variants.
2013年,ACMG,AMP和CAP的成员,代表临床实验室主任和临床医生成立了一个工作组,该工作组依据专家建议、工作组共识和公众反馈开发了一种可以对现有的证据进行加权的系统,并应用此系统对序列变异进行标准分类。为了评估临床实验室的观点,对列入GeneTests.org上位于美国和加拿大的超过100家的测序实验室进行了调研,要求各实验室填写参考术语及变异分类的评估证据。这些实验室有检测包括罕见病、药物基因组学和癌症体细胞突变的经验。第一次调研于2013年2月开展,该调研旨在评估参考术语的偏好,调研结果公布在同年ACMG年会公开论坛上,该年会有超过75个与会者参加。调研结果代表超过45个位于北美的实验室。调研和公开论坛的结果表明: (i) 五级术语系统“致病的”、“可能致病的”、“意义不明确的”、“可能良性的”和“良性的”是优选认可的,且已在多数实验室使用; (ii) 工作组的首要重点应着重于孟德尔疾病和线粒体变异。
In the first survey, laboratories also were asked to provide their protocols for variant assessment, and 11 shared their methods. By analyzing all the protocols submitted, the workgroup developed a set of criteria to weight variant evidence and a set of rules for combining criteria to arrive at one of the five classification tiers. Workgroup members tested the scheme within their laboratories for several weeks using variants already classified in their laboratories and/or by the broader community. In addition, typical examples of variants harboring the most common types of evidence were tested for classification assignment to ensure the system would classify those variants according to current approaches consistently applied by workgroup members. A second survey was sent in August 2013 to the same laboratories identified through GeneTests. org as well as through AMP’s listserv of ~2,000 members, along with the proposed classification scheme and a detailed supplement describing how to use each of the criteria. Laboratories were asked to use the scheme and to provide feedback as to the suitability and relative weighting of each criteria, the ease of use of the classification system, and whether they would adopt such a system in their own laboratory. Responses from over 33 laboratories indicated majority support for the proposed approach, and feedback further guided the development of the proposed standards and guidelines.
在第一次调研中,参与的实验室被要求提供他们的变异评估方法,最终有11个实验室提供并分享了他们的变异评估方法。通过分析所有提交的方法,工作组制定了一组准则,包括变异证据评估的加权标准体系和应用这个标准将变异归类为五类的分类准则。在今后的几周时间里,工作组成员通过在自己实验室或其他机构已进行分类的变异来验证这个方案。另外,还将典型变异的常见证据进行分类,来测试工作组成员达成一致的现有方法是否可以对这些变异进行分类。2013年8月,第二次调研在GeneTests.org上的相同实验室以及AMP清单上的约2000个单位中进行,同时给各单位提供了分类方案和详细的方案补充说明,要求各实验室使用该分类方案并对以下内容进行反馈,包括各标准的适宜性和每个标准的相对权重、分类体系的易用性以及他们是否会在自己的实验室采用这样的体系。来自超过33个实验室的答复表明多数实验室支持所推荐的分类方案,同时,他们的反馈进一步地指导了标准和指南的完善。
In November 2013 the workgroup held a workshop at the AMP meeting with more than 50 attendees, presenting the revised classification criteria and two potential scoring systems. One system is consistent with the approach presented here and the other is a point system whereby each criterion is given a number of points, assigning positive points for pathogenic criteria and negative points for benign criteria, with the total defining the variant class. With an audience-response system, the participants were asked how they would weight each criterion (as strong, moderate or supporting, or not used) during evaluation of variant evidence. Again, the responses were incorporated into the classification system presented here. It should be noted that while the majority of respondents did favor a point system, the workgroup felt that the assignment of specific points for each criterion implied a quantitative level of understanding of each criterion that is currently not supported scientifically and does not take into account the complexity of interpreting genetic evidence.
2013年11月,工作组在AMP会议期间举行了超过50人参加的研讨会,提出了修订后的分类标准和两个评分体系。一个体系与这里介绍的方法是一致的,另一个体系则是一个分数体系,每一项标准都有一个分数,正分数为致病标准,负分数为良性标准,根据总分数进行变异分类。参与者使用此系统并进行反馈,回答在评估变异证据过程中他们如何权衡各个标准(如强、中度或支持、或不使用)。参与者的反馈结果再次综合到这里介绍的分类体系中。但要指出的是,虽然大多数回复更倾向于分数评价体系,但本工作组认为,每个标准中具体分数的设置 量化了对每个标准的理解,但是这一量化指标目前缺乏科学依据,并且没有考虑遗传证据解读时的复杂性。
The workgroup also evaluated the literature for recommendations from other professional societies and working groups that have developed variant classification guidelines for wellstudied genes in breast cancer, colon cancer, and cystic fibrosis and statistical analysis programs for quantitative evaluation of variants in select diseases.While those variant analysis guidelines are useful in a specific setting, it was difficult to apply their proposed criteria to all genes and in different laboratory settings. The variant classification approach described in this article is meant to be applicable to variants in all Mendelian genes, whether identified by single gene tests, multigene panels, exome sequencing, or genome sequencing. We expect that this variant classification approach will evolve as technology and knowledge improve. We should also note that those working in specific disease groups should continue to develop more focused guidance regarding the classification of variants in specific genes given that the applicability and weight assigned to certain criteria may vary by gene and disease.
工作组还评估了文献中推荐的其他专业协会和工作组在乳腺癌、结肠癌和囊性纤维化中已制定的变异分类指南,以及在特定疾病中应用统计分析来进行变异定量评估的方法。这些变异分析指南在特定条件下是有效的,但很难将他们推荐的标准应用于所有基因变异及不同的实验室条件。本文描述的变异分类方法适用于所有孟德尔基因变异,包括单基因、多基因包、外显子组和基因组测序发现的变异。期望这种变异分类方法会随着技术和知识水平的提高而与时俱进。由于不同基因和不同疾病中的应用和加权评估的标准可能不同,特定疾病组的工作应继续,以制定更有针对性的具体基因的变异分类指南。
3.总论
3.1 术语
A mutation is defined as a permanent change in the nucleotide sequence, whereas a polymorphism is defined as a variant with a frequency above 1%. The terms “mutation” and “polymorphism,” however, which have been used widely, often lead to confusion because of incorrect assumptions of pathogenic and benign effects, respectively. Thus, it is recommended that both terms be replaced by the term “variant” with the following modifiers: (i) pathogenic, (ii) likely pathogenic, (iii) uncertain significance, (iv) likely benign, or (v) benign. Although these modifiers may not address all human phenotypes, they comprise a five-tier system of classification for variants relevant to Mendelian disease as addressed in this guidance. It is recommended that all assertions of pathogenicity (including “likely pathogenic”) be reported with respect to a condition and inheritance pattern (e.g., c.1521_1523delCTT (p.Phe508del), pathogenic, cystic fibrosis, autosomal recessive).
突变是指核苷酸序列的永久性改变,而多态性是指频率超过1%的变异。虽然术语“突变”和“多态性”已被广泛使用,但由于这两个术语已经错误地与致病性和良性结果关联了起来,所以往往会造成混淆。因此,建议使用“变异”加以下修饰词替代上述两个术语: 致病性的、可能致病性的、意义不明确的、可能良性的或良性的。虽然这些修饰词不可能适用所有的人类表型,但是正如本指南提出的它包含了孟德尔疾病相关的变异分类五级系统。建议所有致病性(包括可能致病)的结论需要注明疾病及相应的遗传模式(如c.1521_1523delCTT(p.Phe508del),致病性,囊性纤维化,常染色体隐性遗传)。
It should be noted that some laboratories may choose to have additional tiers (e.g., subclassification of variants of uncertain significance, particularly for internal use), and this practice is not considered inconsistent with these recommendations. It should also be noted that the terms recommended here differ somewhat from the current recommendations for classifying copy-number variants detected by cytogenetic microarray.6 The schema recommended for copy-number variants, while also including five tiers, uses “uncertain clinical significance— likely pathogenic” and “uncertain clinical significance—likely benign.” The majority of the workgroup was not supportive of using “uncertain significance” to modify the terms “likely pathogenic” or “likely benign” given that it was felt that the criteria presented here to classify variants into the “likely” categories included stronger evidence than outlined in the copy-number variant guideline and that combining these two categories would create confusion for the health-care providers and individuals receiving clinical reports. However, it was felt that the use of the term “likely” should be restricted to variants where the data support a high likelihood that it is pathogenic or a high likelihood that it is benign. Although there is no quantitative definition of the term “likely,” guidance has been proposed in certain variant classification settings. A survey of the community during an ACMG open forum, however, suggested a much wider range of uses of the term “likely.” Recognizing this, we propose that the terms “likely pathogenic” and “likely benign” be used to mean greater than 90% certainty of a variant either being diseasecausing or benign to provide laboratories with a common, albeit arbitrary, definition. Similarly, the International Agency for Research on Cancer guideline supports a 95% level of certainty of pathogenicity, but the workgroup (confirmed by feedback during the ACMG open forum) felt that clinicians and patients were willing to tolerate a slightly higher chance of error, leading to the 90% decision. It should also be noted that at present most variants do not have data to support a quantitative assignment of variant certainty to any of the five categories given the heterogeneous nature of most diseases. It is hoped that over time experimental and statistical approaches to objectively assign pathogenicity confidence to variants will be developed and that more rigorous approaches to defining what the clinical community desires in terms of confidence will more fully inform terminologies and likelihoods.
应当注意的是,一些实验室可能选择其他等级(如意义不明确的变异的子分类,特别是内部使用时),这种做法不被认为与指南不一致。还应当指出的是,某种程度上本指南推荐的术语与细胞遗传学基因芯片检测的拷贝数变异分类不同。虽然拷贝数变异分类系统也包括五级分类标准,但是它使用“临床意义不明确-可能致病的”和“临床意义不明确-可能良性的”。由于本指南提出的“可能的”变异分类标准比拷贝数变异分类指南中用到的“可能的”包含更强的证据,合并这两个“可能的”分类会使医务工作者和临床报告接收者产生混淆,因此大多数工作组成员不支持使用“意义不明确的”来修饰“可能致病的”或“可能良性的”。然而,有人认为“可能的”一词的使用应限于有数据支持其致病性或良性可能性很大的变异。虽然对“可能的”一词没有量化的定义,但是在某些变异分类系统中已有指导性意见。然而,ACMG开放论坛的一项调查建议“可能的”这一术语具有更广泛的适用性。认识到这一点,建议术语“可能致病的”和“可能良性的”用来说明一个具有大于90%可能引起致病或者可能良性的变异,尽管是人为的界定,但还是给实验室提供了一种共同的定义。类似地,国际癌症机构指南支持致病性的确定水平为95%,但是工作组(通过ACMG公开论坛期间的反馈确认)认为,临床医生和患者愿意容忍略高的错误机会,从而做出确定为90%的决定。还应当指出的是,考虑到多数疾病具有异质性,目前大多数变异没有数据能将它们量化性地归于上述五个变异类别之一。希望随着时间的推移,能够建立实验和统计方法来客观地赋予变异的致病可信度,并且采用更严格的方法来定义临床专业人员所期望达到的可信度,从而能更完整的诠释这些术语及可能性。
The use of new terminologies may require education of the community. Professional societies are encouraged to engage in educating all laboratories as well as health-care providers on the use of these terms, and laboratories also are encouraged to directly educate their ordering physicians.
新术语的使用可能需要专业培训,鼓励专业团队对所有实验室和医务工作者进行这些术语的培训,也鼓励实验室直接对其开具检测报告单的医生进行培训教育。
3.2 命名
A uniform nomenclature, informed by a set of standardized criteria, is recommended to ensure the unambiguous designation of a variant and enable effective sharing and downstream use of genomic information. A standard gene variant nomenclature (http://www.hgvs.org/mutnomen) is maintained and versioned by the Human Genome Variation Society (HGVS), and its use is recommended as the primary guideline for determining variant nomenclature except as noted. Laboratories should note the version being used in their test methods. Tools are available to provide correct HGVS nomenclature for describing variants (https://mutalyzer.nl). Clinical reports should include sequence reference(s) to ensure unambiguous naming of the variant at the DNA level, as well as to provide coding and protein nomenclature to assist in functional interpretations (e.g., “g.” for genomic sequence, “c.” for coding DNA sequence, “p.” for protein, “m.” for mitochondria).
建议通过一套规范的标准对变异进行统一命名来确保变异的明确定义,并实现基因组信息的有效共享和下游使用。标准的基因变异命名由人类基因组变异协会(the Human Genome Variation Society,HGVS)维护和版本化(https://www.hgvs.org/mutnomen),除非另有说明,一般推荐该命名法作为确定变异命名的首要准则。实验室应该注意他们在实验方法中所使用的版本。当描述变异时,可利用这些工具提供正确的HGVS命名(http://mutalyzer.nl)。临床报告应该包含参考序列以确保该变异在DNA水平上的明确命名,并提供编码和蛋白质命名法来协助功能注释(如“g”为基因组序列,“c”为编码DNA序列,“p”为蛋白质,“m”为线粒体)。
The coding nomenclature should be described using the “A” of the ATG translation initiation codon as position number 1. Where historical alternate nomenclature has been used, current nomenclature should be used with an additional notation of the historical naming. The reference sequence should be complete and derived from either the National Center for Biotechnology Information RefSeq database (http://www.ncbi.nlm.nih.gov/RefSeq/) with the version number or the Locus Reference Genomic database (http:// www.lrg-sequence.org). Genomic coordinates should be used and defined according to a standard genome build (e.g., hg19) or a genomic reference sequence that covers the entire gene (including the 5′ and 3′ untranslated regions and promoter). A reference transcript for each gene should be used and provided in the report when describing coding variants. The transcript should represent either the longest known transcript and/or the most clinically relevant transcript. Communitysupported reference transcripts can often be identified through Locus Reference Genomic, the Consensus CDS Database, the Human Gene Mutation Database (http://www.hgmd. cf.ac.uk), ClinVar (http://www.ncbi.nlm.nih.gov/clinvar), or a locus-specific database. However, laboratories should evaluate the impact of the variant on all clinically relevant transcripts, including alternate transcripts that contain additional exons or extended untranslated regions, when there are known variants in these regions that are clinically interpretable.
编码命名应该使用翻译起始密码子ATG中的“A”作为位置编号1来描述。在传统命名已被使用的地方,当今命名应该对传统命名进行额外注释。参考序列应该是完整的,并来源于具有版本号的美国生物技术信息参考序列数据库(http://www.ncbi.nlm.nih.gov/Refseq/)或LRG数据库(http://www.lrg-sequence.org)。基因组坐标应根据标准基因组版本(如hg19)或覆盖整个基因(包括5'和3'非翻译区以及启动子)的基因组参考序列来界定。当描述编码变异时,应该在报告中使用和提供每个基因的一个参考转录本。该转录本应该是最长的已知转录本或者是最具临床相关性的转录本。协会支持的参考转录本通常可以通过LRG数据库(http://www.lrg-sequence.org)、CDS共识数据库(https://www.ncbi.nlm.nih.gov/CCDS/CcdsBrowse.cgi)、人类基因突变数据库(http://www.hgmd.cf.ac.uk)、ClinVar (http://www.ncbi.nlm.nih.gov/clinvar)或特异基因座数据库来确定。然而,当这些区域发生临床可解释的已知变异时,实验室应该评估该变异对所有临床相关的转录本的影响,包括含有其他外显子或非翻译区延伸的可变剪切转录本。
Not all types of variants (e.g., complex variants) are covered by the HGVS recommendations, but possible descriptions for complex variants have been reported. In addition, this ACMG recommendation supports three specific exceptions to the HGVS nomenclature rules: (i) “X” is still considered acceptable for use in reporting nonsense variants in addition to the current HGVS recommendation of “*” and “Ter”; (ii) it is recommended that exons be numbered according to the chosen reference transcript used to designate the variant; and (iii) the term “pathogenic” is recommended instead of “affects function” because clinical interpretation is typically directly evaluating pathogenicity.
HGVS(https://www.hgvs.org/mutnomen)并未覆盖所有类型的变异(如复杂变异),但是复杂变异的可能描述已被报道。此外,ACMG支持HGVS命名规则之外的三种特殊例外: (i) 除了当今HGVS推荐的“*”和“Ter”,“X”仍然被认为用于报告无义变异; (ii) 建议根据指定变异选择的参考转录本对外显子进行编号; (iii) 通常因为临床解释直接评估致病性,所以推荐使用术语“致病性”而不是“影响功能”。
3.3 文献及数据库使用
A large number of databases contain a growing number of variants that are continuously being discovered in the human genome. When classifying and reporting a variant, clinical laboratories may find valuable information in databases, as well as in the published literature. As noted above, sequence databases can also be used to identify appropriate reference sequences. Databases can be useful for gathering information but should be used with caution.
目前人类基因组中大量变异不断被发现,且已被许多数据库广泛收录。当临床实验室需要对某一变异进行分类并出具报告时,可在已有的数据库及发表的文献中寻找到有价值的参考信息。如上文提及,序列数据库还可用于确定合适的参考序列。数据库有助于信息收集,但需谨慎使用。
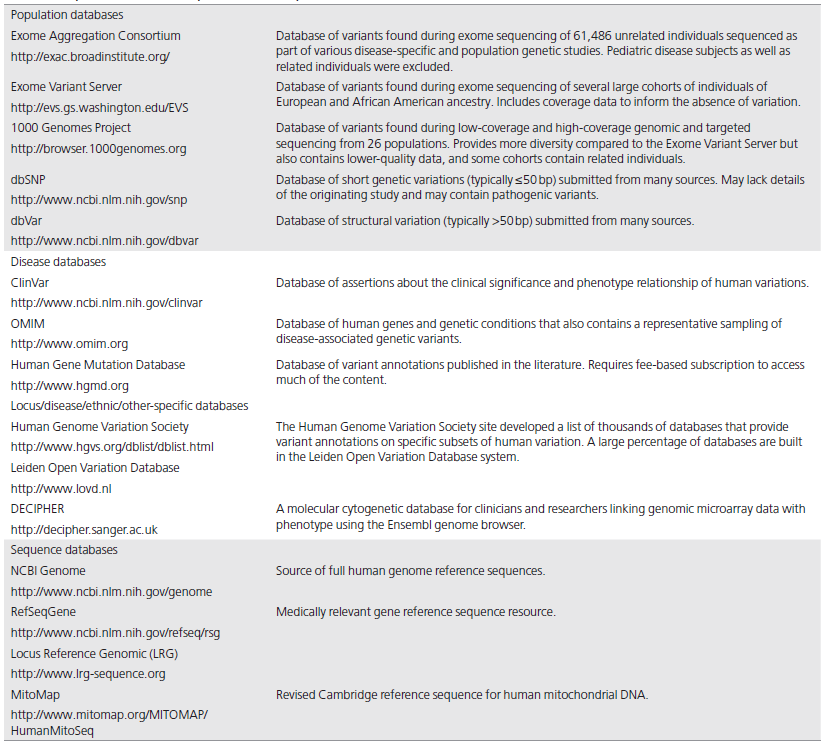
Population databases (Table 1) are useful in obtaining the frequencies of variants in large populations. Population databases cannot be assumed to include only healthy individuals and are known to contain pathogenic variants. These population databases do not contain extensive information regarding the functional effect of these variants or any possible associated phenotypes. When using population databases, one must determine whether healthy or disease cohorts were used and, if possible, whether more than one individual in a family was included, as well as the age range of the subjects.
人群数据库(表1)适用于获取某变异在大规模人群中发生频率的相关信息。需要注意的是,人群数据库中的信息不仅来源于健康个体,也包含致病性的变异。另外,人群数据库并不包含变异的功能效应及可能关联的表型等相关信息。在使用人群数据库时,须明确数据库收录的是健康群体的信息还是患病群体的信息; (如能确认)数据库是否收录了同一家庭多名成员的信息以及数据库收录的受试者的年龄范围。
Disease databases (Table 1) primarily contain variants found in patients with disease and assessment of the variants’ pathogenicity. Disease and gene-specific databases often contain variants that are incorrectly classified, including incorrect claims published in the peer-reviewed literature, because many databases do not perform a primary review of evidence. When using disease databases, it is important to consider how patients were ascertained, as described below.
疾病数据库(表1)主要包含病患中发现的变异以及对其致病性的评估。疾病数据库和特定基因的数据库常包含一些分类错误的变异,这些变异在已发表的同行评审的文献中被错误判定,而多数数据库在收录变异相关信息时并未对证据进行基本的审核。因此,在使用疾病数据库时,考虑患者是如何被确诊的尤为重要,如下所述:
When using databases, clinical laboratories should (i) determine how frequently the database is updated, whether data curation is supported, and what methods were used for curation;(ii) confirm the use of HGVS nomenclature and determine the genome build and transcript references used for naming variants; (iii) determine the degree to which data are validated for analytical accuracy (e.g., low-pass nextgeneration sequencing versus Sanger-validated variants) and evaluate any quality metrics that are provided to assess data accuracy, which may require reading associated publications; and (iv) determine the source and independence of the observations listed.
当使用数据库时,临床实验室应做到: (i) 确定数据库的更新频率,确定数据库收录相关数据时是否进行了校勘,以及采用什么方法进行数据校勘; (ii) 确认采用HGVS命名体系,并确定描述变异的基因组版本和转录本参考序列; (iii) 确定数据分析准确度的验证程度(如变异是源自于低覆盖的新一代测序,还是通过了Sanger测序验证),并分析用于评估数据准确度的各种指标,要获得这些信息可能需要阅读相关的文献; (iv) 确定收录对象的来源及其唯一性。
Variant assessment also includes searching the scientific and medical literature. Literature using older nomenclature and classification or based on a single observation should be used with caution. When identifying individuals and families with a variant, along with associated phenotypes, it is important to consider how patients were ascertained. This caveat is important when assessing data from publications because affected individuals and related individuals are often reported multiple times, depending on the context and size of the study. This may be due to authorship overlap, interlaboratory collaborations, or a proband and family members being followed across different clinical systems. This may mistakenly lead to duplicate counting of affected patients and a false increase in variant frequency. Overlapping authorship or institutions is the first clue to the potential for overlapping data sets.
变异解读也需要检索科学和医学文献。在参考一些采用旧的命名和分类系统或基于单一观察结果的文献时需要慎重。在参考携带某一变异并伴有相关表型的个体和家系的信息时,考虑患者是如何被确诊尤为重要。在评估这些文献的数据时需要谨慎客观,这是由于受累患者及相关个体在基于不同背景和规模的研究中常常被多次重复报道。重复报道的发生可能是由于作者重叠、实验室间合作或先证者及其家庭成员同时被不同临床系统随访。而这些重复报道可能会导致受累个体被错误地重复计数,进而使变异频率假性增高。作者或其研究机构互相重叠是发现数据集重复的第一线索。
Clinical laboratories should implement an internal system to track all sequence variants identified in each gene and clinical assertions when reported. This is important for tracking genotype–phenotype correlations and the frequency of variants in affected and normal populations. Clinical laboratories are encouraged to contribute to variant databases, such as ClinVar, including clinical assertions and evidence used for the variant classification, to aid in the continued understanding of the impact of human variation. Whenever possible, clinical information should be provided following Health Insurance Portability and Accountability Act regulations for privacy. Clinical laboratories are encouraged to form collaborations with clinicians to provide clinical information to better understand how genotype influences clinical phenotype and to resolve differences in variant interpretation between laboratories. Because of the great potential to aid clinical laboratory practice, efforts are underway for clinical variant databases to be expanded and standardized. Standardization will provide easier access to updated information as well as facilitate submission from the clinical laboratory. For example, the ClinVar database allows for the deposition of variants with clinical observations and assertions, with review status tracked to enable a more transparent view of the levels of quality of the curation.
临床实验室应建立一个内部系统对已报告的基因序列变异及临床诊断进行记录。这对于分析基因型-表型之间的相关性,以及该变异在患者和正常人群中的发生频率尤为重要。临床实验室也应该积极提交变异数据到相关数据库,如ClinVar数据库,包含提交临床评估信息以及用于变异分类的证据,以帮助人们不断加深对人类遗传变异所产生的效应的理解。在任何时候,提供临床数据应遵循“健康保险携带和责任法案 (HIPAA)”对个人隐私保护的规定。临床实验室应与临床医生合作,以获得临床信息,从而更好地理解基因型是如何影响临床表型的,并解决不同实验室对遗传变异解读存在差异的问题。临床变异数据库极大地促进临床实验室工作的开展,因此需对其进行扩展并标准化。标准化便于临床实验室获取数据库的最新信息,同时有助于提交更新的信息。例如,ClinVar数据库允许变异连同临床表型和诊断相关信息一并提交,同时追踪提交变异的审核状态,以便对校勘质量的水平提供一个更加透明的概貌。
3.4 生物信息学计算预测程序
A variety of in silico tools, both publicly and commercially available, can aid in the interpretation of sequence variants. The algorithms used by each tool may differ but can include determination of the effect of the sequence variant at the nucleotide and amino acid level, including determination of the effect of the variant on the primary and alternative gene transcripts, other genomic elements, as well as the potential impact of the variant on the protein. The two main categories of such tools include those that predict whether a missense change is damaging to the resultant protein function or structure and those that predict whether there is an effect on splicing (Table 2). Newer tools are beginning to address additional noncoding sequences.
各种公共和商业化计算机工具可以辅助解读序列变异。每种工具使用的算法可能有差异,但都会包含序列变异在核苷酸及氨基酸水平上作用影响的判断,包括变异对主要转录本,可变转录本,其他基因组元件影响作用的确认,也包括对蛋白质潜在影响作用的判定。这些工具主要分为两类: 一类可以预测错义变异是否会破坏蛋白质的功能或结构; 另一种可以预测是否影响剪接(表2)。新的工具已可以处理额外的非编码序列。
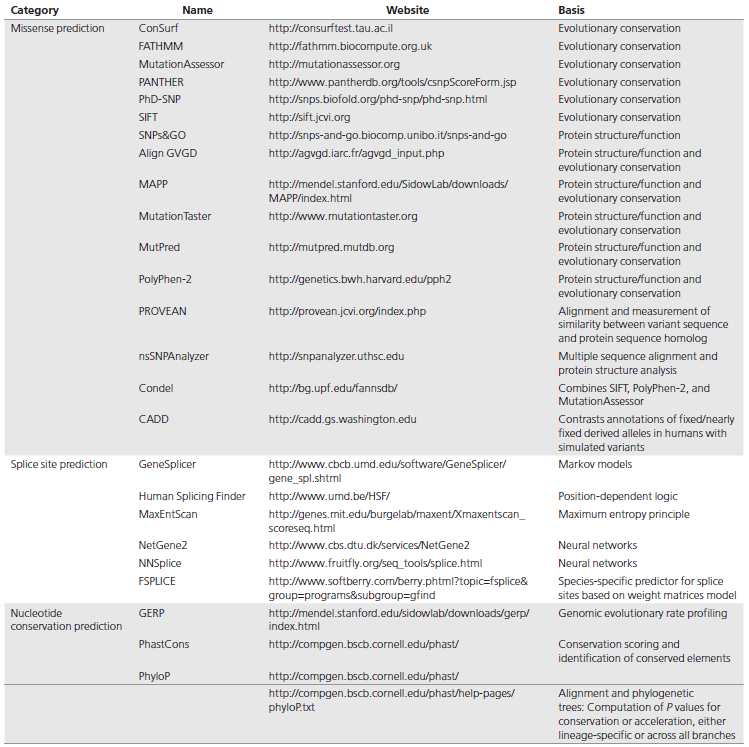
The impact of a missense change depends on criteria such as the evolutionary conservation of an amino acid or nucleotide, the location and context within the protein sequence, and the biochemical consequence of the amino acid substitution. The measurement of one or a combination of these criteria is used in various in silico algorithms that assess the predicted impact of a missense change. Several efforts have evaluated the performance of available prediction software to compare them with each other and to assess their ability to predict “known” disease-causing variants. In general, most algorithms for missense variant prediction are 65–80% accurate when examining known disease variants. Most tools also tend to have low specificity, resulting in overprediction of missense changes as deleterious, and are not as reliable at predicting missense variants with a milder effect.18 The in silico tools more commonly used for missense variant interpretation in clinical laboratories include PolyPhen2, SIFT, and MutationTaster. A list of in silico tools used to predict missense variants can be found in Table 2.
错义改变的影响作用是由不同的条件决定的,例如一个氨基酸或核苷酸的进化保守性、其在蛋白质序列中的位置及其上下游序列,以及氨基酸置换导致的生化结果等。对各种计算机算法中的一个或几个条件进行评测可以进一步评估错义改变带来的影响。已经有一些工作在评估预测软件的预测性能,是通过对这些预测软件之间的相互比较评估他们预测已知致病突变的能力来实现的。一般情况下,多数算法预测已知致病的错义突变的准确率能达到65%~80%。但是大多数工具的特异性较低,导致有些错义改变被过度预测为有害突变,而且对于影响较小的错义变异的预测也不可靠。目前临床实验室常用的错义变异解读工具有PolyPhen 2,SIFT和MutationTaster。用于预测错义变异的生物信息分析工具见表2。
Multiple software programs have been developed to predict splicing as it relates to the creation or loss of splice sites at the exonic or intronic level. In general, splice site prediction tools have higher sensitivity (~90–100%) relative to specificity (~60–80%) in predicting splice site abnormalities. Some of the in silico tools commonly used for splice site variant interpretation are listed in Table 2.
目前已开发出许多用于预测剪接的软件,这是基于内含子或外显子水平上剪接位点的丢失或产生原理基础上而完成的。一般情况下,相对于特异性(60%~80%),预测工具在预测剪接位点异常方面具有较高的敏感性(~90%~100%)。一些常用的剪接位点预测分析计算工具见表2。
While many of the different software programs use different algorithms for their predictions, they have similarities in their underlying basis; therefore, predictions combined from different in silico tools are considered as a single piece of evidence in sequence interpretation as opposed to independent pieces of evidence. The use of multiple software programs for sequence variant interpretation is also recommended because the different programs each have their own strengths and weaknesses, depending on the algorithm; in many cases performance can vary by the gene and protein sequence. These are only predictions, however, and their use in sequence variant interpretation should be implemented carefully. It is not recommended that these predictions be used as the sole source of evidence to make a clinical assertion.
虽然许多不同的分析软件程序使用不同的算法进行预测,但其基本原理是相似的; 因此,在序列解读中,不同软件工具组合的预测结果被视为单一证据而不是相互独立的证据。因为每个软件工具基于他们使用的算法都各有优缺点,所以仍然建议使用多种软件进行序列变异解读; 很多情况下,预测性可能因为基因和蛋白质序列的不同而有差异。无论如何,这些软件分析结果只是预测,他们在序列变异解读中的应用应该慎重。不建议仅使用这些预测结果作为唯一证据来源进行临床判断。
4. 序列变异解读的拟定标准
The following approach to evaluating evidence for a variant is intended for interpretation of variants observed in patients with suspected inherited (primarily Mendelian) disorders in a clinical diagnostic laboratory setting. It is not intended for the interpretation of somatic variation, pharmacogenomic (PGx) variants, or variants in genes associated with multigenic non- Mendelian complex disorders. Care must be taken when applying these rules to candidate genes (“genes of uncertain significance” (GUS)) in the context of exome or genome studies (see Special Considerations below) because this guidance is not intended to fulfill the needs of the research community in its effort to identify new genes in disease.
以下评估变异证据的方法是用了解释在临床诊断实验室中具有疑似遗传(主要指孟德尔遗传)疾病患者的变异。并不适用于解读体细胞变异、药物基因组(PGx)变异、或者是多基因非孟德尔复杂疾病相关的基因变异。在外显子组或基因组研究中,对候选基因(意义不明确的基因(GUS))应用这些准则时应当谨慎(见下面注意事项),因为本指南目的不是满足鉴定新致病基因的研究需求。
Although these approaches can be used for evaluating variants found in healthy individuals or secondary to the indication for testing, further caution must be used, as noted in several parts of the guideline, given the low prior likelihood that most variants unrelated to the indication are pathogenic. Although we expect that, in general, these guidelines will apply for variant classification regardless of whether the variant was identified through analysis of a single gene, gene panel, exome, genome, or transcriptome, it is important to consider the differences between implicating a variant as pathogenic (i.e., causative) for a disease and a variant that may be predicted to be disruptive/ damaging to the protein for which it codes, but is not necessarily implicated in a disease. These rules are intended to determine whether a variant in a gene with a definitive role in a Mendelian disorder may be pathogenic for that disorder. Pathogenicity determination should be independent of interpreting the cause of disease in a given patient. For example, a variant should not be reported as pathogenic in one case and not pathogenic in another simply because the variant is not thought to explain disease in a given case. Pathogenicity should be determined by the entire body of evidence in aggregate, including all cases studied, arriving at a single conclusion.
虽然这些方法可用于评估在健康个体中发现的变异或与测试指征不相关的变异,但是正如在指南的几个部分中所述,对于与指征无关的有较低先验致病性的变异时需更加谨慎。尽管期望本指南适用于变异分类,无论其是通过分析单基因,基因包,外显子组,基因组或者转录组而鉴定的,重要的是要关注与疾病有关的致病变异和虽然预测为对蛋白有破坏/损伤但却与疾病无充分关联的变异之间的区别。这些规则旨在确定在孟德尔遗传病中有明确作用的基因的变异是否对该遗传疾病是致病的。针对具体的病人,致病性判定应该独立于对疾病病因的解读。例如,某变异在一个案例中被评估为“致病的”,而在另一个案例中,由于不能解释该疾病,就对这个位点不给出“致病的”评价,这样的情况是绝对不允许的。确定致病性需要将全部的证据汇集在一起,包括所有的案例分析,最终得出一个结论。
This classification approach may be somewhat more stringent than laboratories have applied to date. They may result in a larger proportion of variants being categorized as uncertain significance. It is hoped that this approach will reduce the substantial number of variants being reported as “causative” of disease without having sufficient supporting evidence for that classification. It is important to keep in mind that when a clinical laboratory reports a variant as pathogenic, health-care providers are highly likely to take that as “actionable” and to alter the treatment or surveillance of a patient or remove such management in a genotype-negative family member, based on that determination (see How Should Health-Care Providers Use These Guidelines and Recommendations, below).
此指南的分类方法可能比目前实验室应用的标准更为严格。这将导致很大一部分的变异被归类为“意义不明确的”。希望这种方法可以大量减少那些没有足够分类证据支持而报告为致病原因的变异。需要注意的是,当临床实验室报告一个变异为“致病的”时,医疗单位很可能把其当作“可指导临床作为的(actionable)”,基于这个判断,从而会改变对患者的治疗、监测,或去除对基因型为阴性的家庭成员的治疗、监测(参见下面的医务工作者应该如何使用这些指南和建议)。
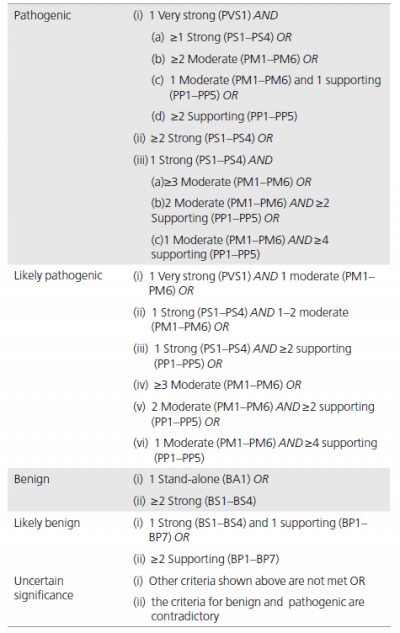
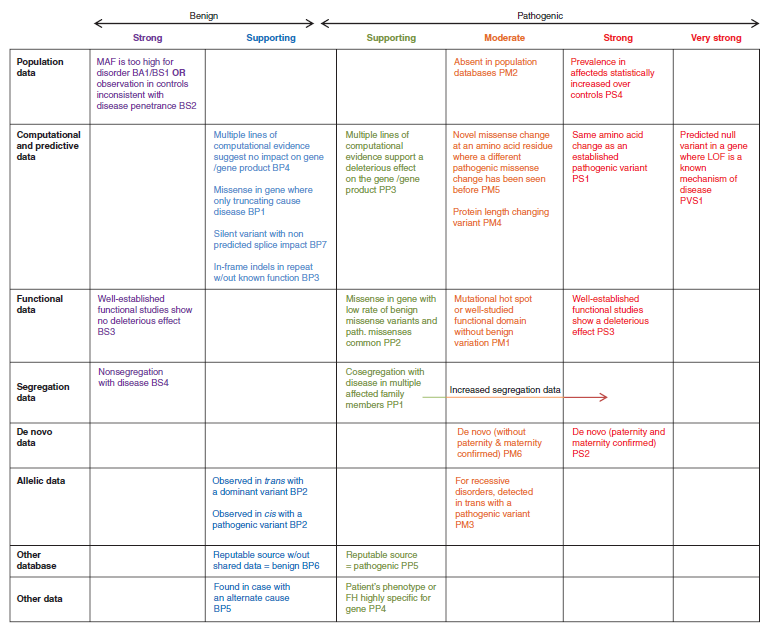
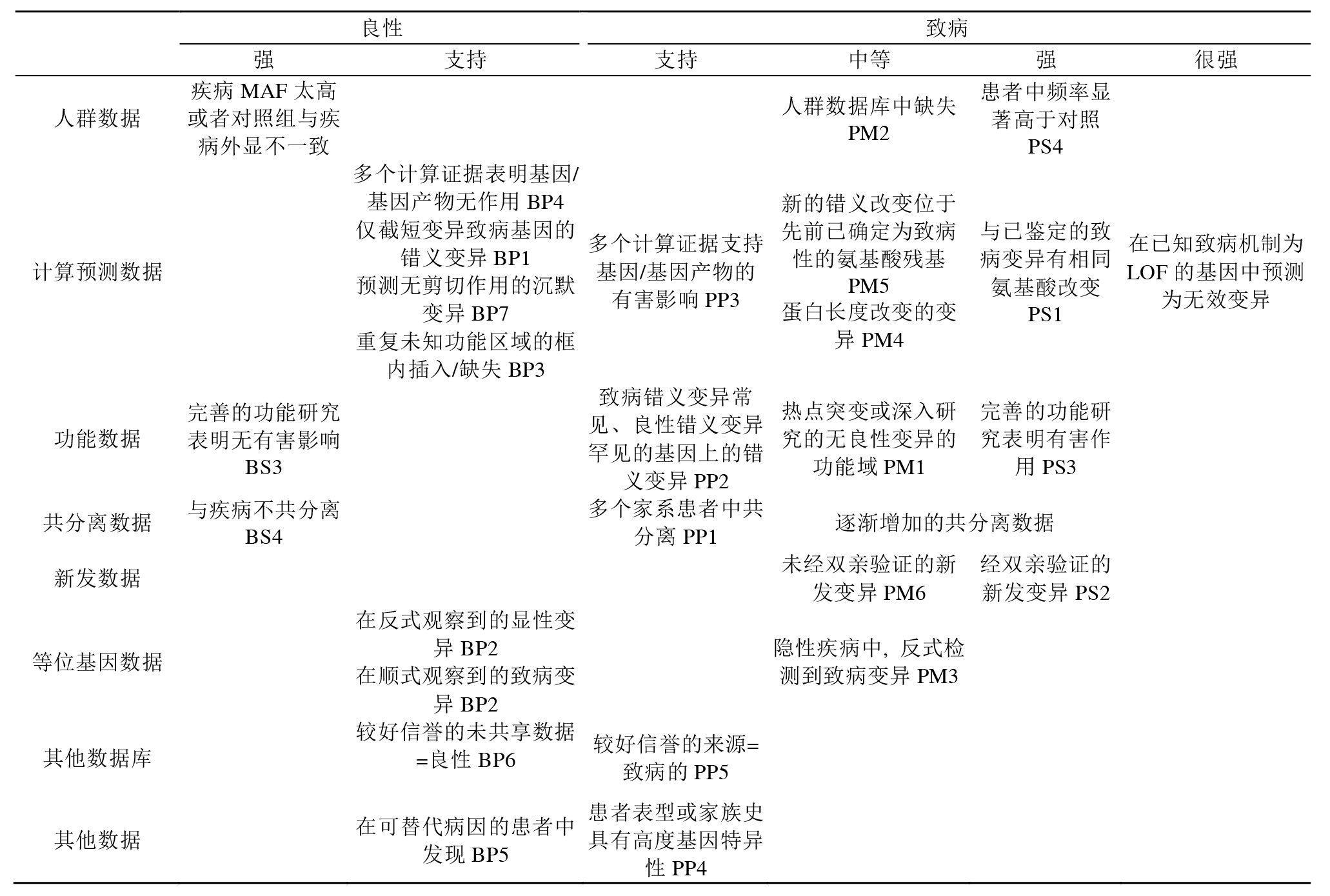
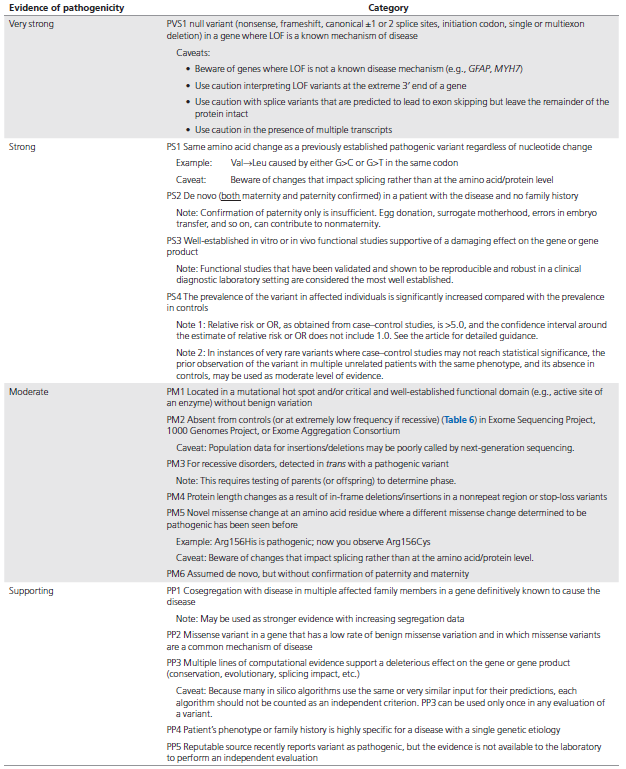
We have provided two sets of criteria: one for classification of pathogenic or likely pathogenic variants (Table 3) and one for classification of benign or likely benign variants (Table 4). Each pathogenic criterion is weighted as very strong (PVS1), strong (PS1–4); moderate (PM1–6), or supporting (PP1–5), and each benign criterion is weighted as stand-alone (BA1), strong (BS1– 4), or supporting (BP1–6). The numbering within each category does not convey any differences of weight and is merely labeled to help refer to the different criteria. For a given variant, the user selects the criteria based on the evidence observed for the variant. The criteria then are combined according to the scoring rules in Table 5 to choose a classification from the five-tier system. The rules apply to all available data on a variant, whether gathered from the current case under investigation or from well-vetted previously published data. Unpublished case data may also be obtained through public resources (e.g., ClinVar or locus specific databases) and from a laboratory’s own database. To provide critical flexibility to variant classification, some criteria listed as one weight can be moved to another weight using professional judgment, depending on the evidence collected. For example, rule PM3 could be upgraded to strong if there were multiple observations of detection of the variant in trans (on opposite chromosomes) with other pathogenic variants (see PM3 BP2 cis/trans Testing for further guidance). By contrast, in situations when the data are not as strong as described, judgment can be used to consider the evidence as fulfilling a lower level (e.g., see PS4, Note 2 in Table 3). If a variant does not fulfill criteria using either of these sets (pathogenic or benign), or the evidence for benign and pathogenic is conflicting, the variant defaults to uncertain significance. The criteria, organized by type and strength, is shown in Figure 1. Please note that expert judgment must be applied when evaluating the full body of evidence to account for differences in the strength of variant evidence.
本指南提供了两套标准: 一是用于对致病或可能致病的变异进行分类(表3),另一是用于对良性或可能良性的变异进行分类(表4)。致病变异标准可分为非常强(very strong,PVS1),强(strong,PS1~4); 中等(moderate,PM1~6),或辅助证据(supporting,PP1~5)。良性变异证据可分为独立(stand-alone,BA1),强(strong,BS1~4),或辅助证据(BP1~6)。其中,数字只是作为有助于参考的分类标注,不具有任何意义。每个类别中的数字不表示分类的任何差异,仅用来标记以帮助指代不同的规则。对于一个给定的变异,用户基于观察到的证据来选择标准。根据表5的评分规则把标准组合起来进而从5级系统中选择一个分类。这些规则适用于变异上的所有可用数据,无论是基于调查现有案例获得的数据,还是来源于先前公布的数据。未发表的数据也可以通过公共数据库(如ClinVar或位点特异数据库)和实验室自有数据库获得。为了对变异分类具有较好灵活性,基于收集的证据和专业判断,可以把某些依据用到不同的证据水平上去。例如,如果一个变异多次和已知致病性变异处于反式位置(位于另一染色体上),PM3可以上调到强(进一步指导见PM3 BP2顺/反式检测)。相反,在数据并不像描述的那么强的情况下,可以改判变异到一个较低的水平(见表3注2 PS4)。如果一个变异不符合分类标准(致病的或良性的),或良性和致病的证据是相互矛盾的,则默认该变异为“意义不确定的”。程度判断评价标准如图1所示。请注意,当考虑所有依据以解读变异证据强度的差异时,须专家介入进行判断。
The following is provided to more thoroughly explain certain concepts noted in the criteria for variant classification (Tables 3 and 4) and to provide examples and/or caveats or pitfalls in their use. This section should be read in concert with Tables 3 and 4.
下面提供更详细的变异分类标准(表3和4)中提及的某些概念的解释,并提供实际使用中的实例和/或误区或易犯错误的地方。这部分应该与表3及4一同阅读。
4.1 PVS1 极强致病性变异
Certain types of variants (e.g., nonsense, frameshift, canonical ±1 or 2 splice sites, initiation codon, single exon or multiexon deletion) can often be assumed to disrupt gene function by leading to a complete absence of the gene product by lack of transcription or nonsense-mediated decay of an altered transcript. One must, however, exercise caution when classifying these variants as pathogenic by considering the following principles:
某些特定类型的变异(如无义突变、移码突变、经典剪接位点±1或2点突变、起始密码子变异、单个或多个外显子缺失)被认为因无转录产物或由无义突变引起的转录子降解,导致基因产物完全缺失而破坏基因功能。当将这类变异归类为致病性时,从业人员需谨慎考虑以下原则:
(i) When classifying such variants as pathogenic, one must ensure that null variants are a known mechanism of pathogenicity consistent with the established inheritance pattern for the disease. For example, there are genes for which only heterozygous missense variants cause disease and null variants are benign in a heterozygous state (e.g., many hypertrophic cardiomyopathy genes). A novel heterozygous nonsense variant in the MYH7 gene would not be considered pathogenic for dominant hypertrophic cardiomyopathy based solely on this evidence, whereas a novel heterozygous nonsense variant in the CFTR gene would likely be considered a recessive pathogenic variant.
(i) 当将该类变异归类为致病性时,需确认无功能变异(null variants)是已知的致病机理,且与该疾病的遗传模式相一致。例如,有些基因(如许多肥厚性心肌病基因)只有杂合错义突变时才致病,而杂合无功能变异却是良性的。仅基于这一项证据来看,对显性肥厚性心肌病来说,MYH7基因上出现一个新的杂合无义突变不一定是致病的,而CFTR基因上出现一个新的杂合无义突变则有可能是一个隐性致病变异。
(ii) One must also be cautious when interpreting truncating variants downstream of the most 3′ truncating variant established as pathogenic in the literature. This is especially true if the predicted stop codon occurs in the last exon or in the last 50 base pairs of the penultimate exon, such that nonsense-mediated decay would not be predicted, and there is a higher likelihood of an expressed protein. The length of the predicted truncated protein would also factor into the pathogenicity assignment, however, and such variants cannot be interpreted without a functional assay.
(ii) 当文献中将3′远端下游截短变异注释成致病突变时,要特别小心。特别是当所预测的终止密码子出现在最后一个外显子,或者出现在倒数第二个外显子的最后50个碱基对时,这种无义突变介导的转录降解可能不会发生,这个蛋白很可能会表达。据此所预测的截短蛋白的长度也是致病性评估的因素,但这些变异未经功能分析是无法进行判定的。
(iii) For splice-site variants, the variant may lead to exon skipping, shortening, or inclusion of intronic material as a result of alternative donor/acceptor site usage or creation of new sites. Although splice-site variants are predicted to lead to a null effect, confirmation of impact requires functional analysis by either RNA or protein analysis. One must also consider the possibility of an in-frame deletion/insertion, which could retain the critical domains of the protein and hence lead to either a mild or neutral effect with a minor length change (PM4) or a gain-of-function effect.
(iii) 就剪接位点变异而言,因外显子剪切位点的供体/受体位点改变或产生了新的剪切位点,从而可能导致外显子丢失、缩短,也可能会使内含子序列变成外显子部分。虽然剪切位点变异可能被预测为无功能变异,然而该变异类型造成的影响需要通过RNA或蛋白质功能分析确认。还必须考虑可读框内缺失/插入的可能性,其长度变化较小(PM4),可以保留蛋白质的关键结构域,因此导致轻微或中性效应,或功能获得效应。
(iv) Considering the presence of alternate gene transcripts and understanding which are biologically relevant, and in which tissues the products are expressed, are important. If a truncating variant is confined to only one or not all transcripts, one must be cautious about overinterpreting variant impact given the presence of the other protein isoforms.
(iv) 基因会有不同的转录本,哪一种转录本与生物学功能相关,在哪些组织会表达哪些转录本,这些都是需要进行重点考虑的。如果一个截短变异只限于一个或并非所有转录本,则必须谨慎考虑到可能存在其他同功型蛋白质,防止过度解释。
(v) One must also be cautious in assuming that a null variant will lead to disease if found in an exon where no other pathogenic variants have been described, given the possibility that the exon may be alternatively spliced. This is particularly true if the predicted truncating variant is identified as an incidental finding (unrelated to the indication for testing), given the low prior likelihood of finding a pathogenic variant in that setting.
(v) 如果发现一个无功能变异位于某个外显子上,而该外显子先前无致病变异报道,那么该外显子可能被选择性剪切了,此时需要谨慎考虑该变异的致病性。当预测的截短变异是偶然发现时(与检测指征无关)则应特别小心,在这种情况下该位点致病的可能性非常低。
4.2 PS1 突变为同一氨基酸
In most cases, when one missense variant is known to be pathogenic, a different nucleotide change that results in the same amino acid (e.g., c.34G>C (p.Val12Leu) and c.34G>T (p.Val12Leu)) can also be assumed to be pathogenic, particularly if the mechanism of pathogenicity occurs through altered protein function. However, it is important to assess the possibility that the variant may act directly through the specific DNA change (e.g., through splicing disruption as assessed by at least computational analysis) instead of through the amino acid change, in which case the assumption of pathogenicity may no longer be valid.
多数情况下,尤其是当致病机制是蛋白质功能发生改变时,如已确定某一错义变异是致病变异,应考虑到与其位于同一变异位点的不同形式的碱基改变也可能产生相同的错义突变结果——氨基酸改变相同(如c.34G>C(p.Val12Leu)和c.34G>T(p.Val12Leu)),那么,这些变异也应是致病突变。此外,还应考虑到,变异可能不是通过改变氨基酸的水平,而是通过改变DNA的序列来发挥作用,例如,破坏剪接位点(可通过软件分析确定),在这种情况下,上述的假设是不成立的。
4.3 PS2 PM6 新发变异
A variant observed to have arisen de novo (parental samples testing negative) is considered strong support for pathogenicity if the following conditions are met: (i) Both parental samples were shown through identity testing to be from the biological parents of the patient. Note that PM6 applies if identity is assumed but not confirmed. (ii) The patient has a family history of disease that is consistent with de novo inheritance (e.g., unaffected parents for a dominant disorder). It is possible, however, that more than one sibling may be affected because of germ-line mosaicism. (iii) The phenotype in the patient matches the gene’s disease association with reasonable specificity. For example, this argument is strong for a patient with a de novo variant in the NIPBL gene who has distinctive facial features, hirsutism, and upper-limb defects (i.e., Cornelia de Lange syndrome), whereas it would be weaker for a de novo variant found by exome sequencing in a child with nonspecific features such as developmental delay.
当我们将一个新发变异(父母样本检测结果阴性)归类为强的致病证据时,需要满足以下条件: (i) 身份检验表明患者的父母是其生物学父母。注意如果父母的身份是假定的而没有被证实,则判定为PM6; (ii) 患者的家族史符合新发变异特征。例如,显性遗传病患者的父母均未患病。在存在生殖细胞嵌合现象时也可能有1个以上同胞患病; (iii) 患者的表型与变异基因异常引起的表型相吻合。例如,患者具有特殊面容、多毛和上肢缺陷(即Cornelia de Lange综合征),检测到NIPBL基因的新生突变即为强致病证据,而患者仅表现为非特异性的发育迟缓,通过外显子组测序发现的该基因的新发变异,则判断此变异致病性的证据较弱。
4.4 PS3 BS3 功能研究
Functional studies can be a powerful tool in support of pathogenicity; however, not all functional studies are effective in predicting an impact on a gene or protein function. For example, certain enzymatic assays offer well-established approaches to assess the impact of a missense variant on enzymatic function in a metabolic pathway (e.g., α-galactosidase enzyme function). On the other hand, some functional assays may be less consistent predictors of the effect of variants on protein function. To assess the validity of a functional assay, one must consider how closely the functional assay reflects the biological environment. For example, assaying enzymatic function directly from biopsied tissue from the patient or an animal model provides stronger evidence than expressing the protein in vitro. Likewise, evidence is stronger if the assay reflects the full biological function of the protein (e.g., substrate breakdown by an enzyme) compared with only one component of function (e.g., adenosine triphosphate hydrolysis for a protein with additional binding properties). Validation, reproducibility, and robustness data that assess the analytical performance of the assay and account for specimen integrity, which can be affected by the method and time of acquisition, as well as storage and transport, are important factors to consider. These factors are mitigated in the case of an assay in a Clinical Laboratory Improvement Amendments laboratory–developed test or commercially available kit. Assays that assess the impact of variants at the messenger RNA level can be highly informative when evaluating the effects of variants at splice junctions and within coding sequences and untranslated regions, as well as deeper intronic regions (e.g., messenger RNA stability, processing, or translation). Technical approaches include direct analysis of RNA and/or complementary DNA derivatives and in vitro minigene splicing assays.
功能实验研究是一种研究变异致病性的非常强大的工具,然而并非所有的功能研究都能有效地预测基因或蛋白的功能。例如,一些酶学实验利用成熟完善的方法可以用来评估错义变异在代谢途径中对酶活性的影响(如α-半乳糖苷酶功能实验); 而另一方面,某些功能实验在评估变异对蛋白质功能的影响时缺乏一致性。评估一个功能检测方法是否有效时,必须考虑该功能实验多大程度上反映了其发挥功能的生物环境。例如,与体外表达蛋白相比,直接在患者或动物模型的活检组织中进行酶的功能实验更有说服力。同样,可以反映蛋白质全部生物学功能(如酶分解底物功能)的实验则比仅反映一部分功能(如一种有附带结合能力的蛋白水解ATP的功能)的实验证据性更强。功能实验的有效性、重复性和稳定性应重点考虑,这些参数用来评估功能实验的分析性能以及判定样本诊断信息的完整性,该完整性容易受标本采集的方法及时间、存储及运输的影响。CLIA(临床实验室改进修正案)认证实验室建立的检测方法或商品化试剂盒可减少这些因素对实验的影响。评估变异在剪接位点、编码序列、非翻译区以及更深的内含子区域的影响时,对变异在信使RNA水平(如信使RNA的稳定性、加工或翻译)进行评估,可以提供丰富的信息。相关的技术方法包括对RNA和/或互补DNA衍生物进行直接分析,以及体外微小基因剪接分析。
4.5 PS4 PM2 BA1 BS1 BS2 变异频率及对照人群的使用
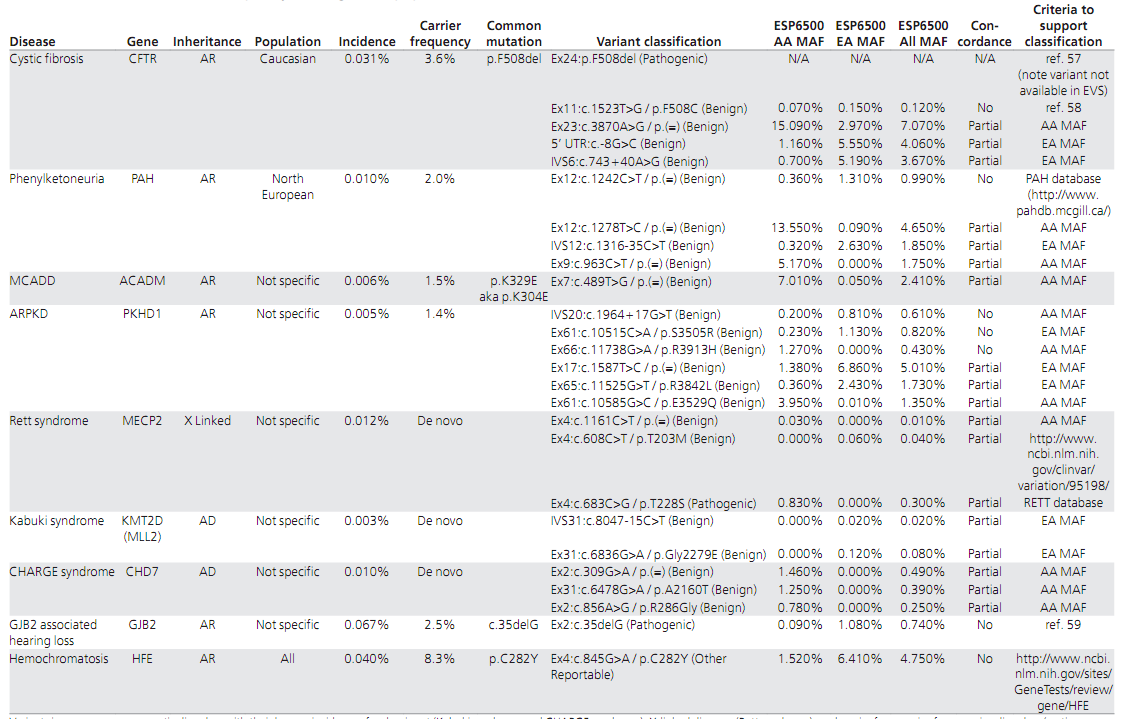
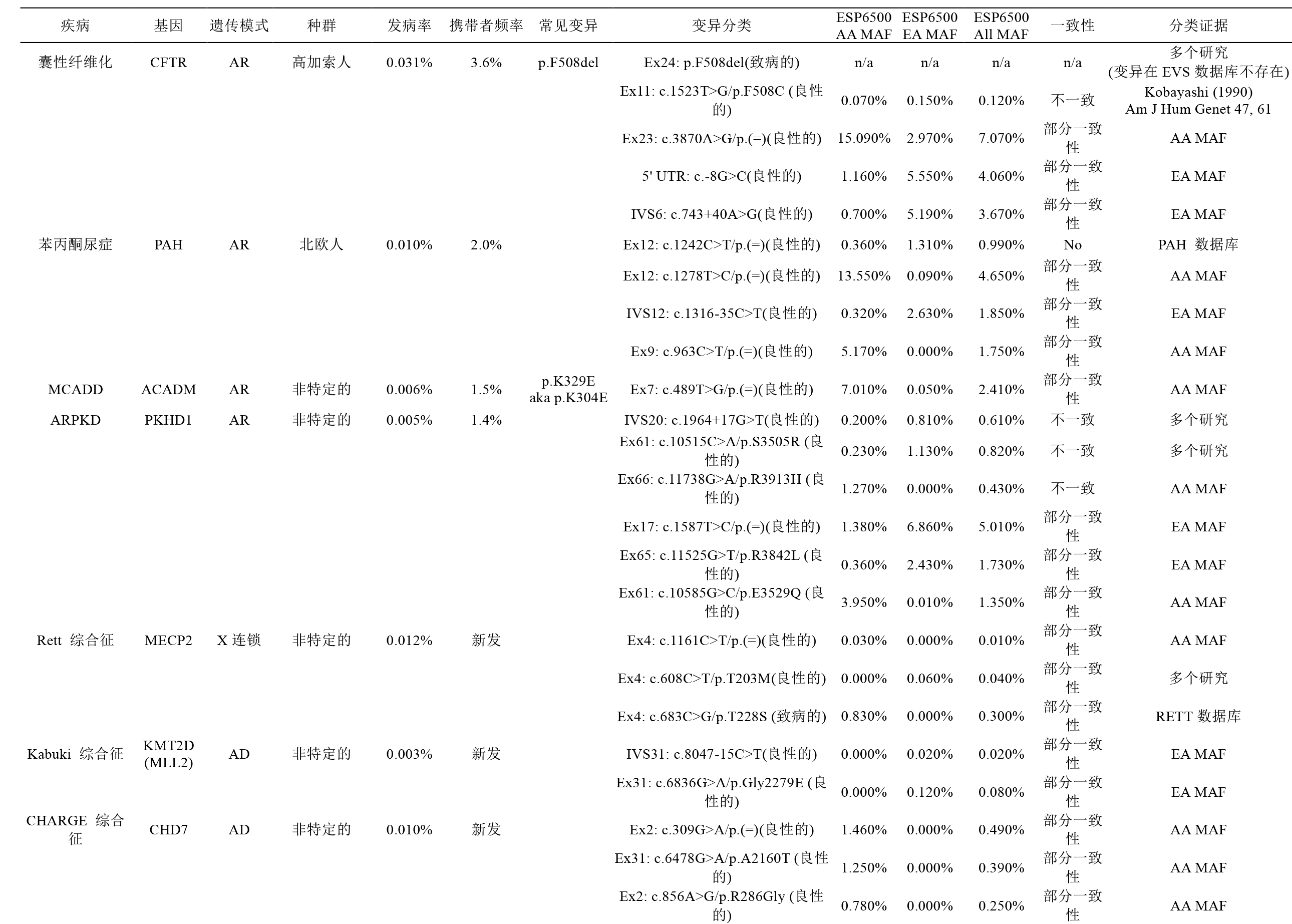
Assessing the frequency of a variant in a control or general population is useful in assessing its potential pathogenicity. This can be accomplished by searching publicly available population databases (e.g., 1000 Genomes Project, National Heart, Lung, and Blood Institute Exome Sequencing Project Exome Variant Server, Exome Aggregation Consortium; Table 1), as well as using race-matched control data that often are published in the literature. The Exome Sequencing Project data set is useful for Caucasian and African American populations and has coverage data to determine whether a variant is absent. Although the 1000 Genomes Project data cannot be used to assess the absence of a variant, it has a broader representation of different racial populations. The Exome Aggregation Consortium more recently released allele frequency data from >60,000 exomes from a diverse set of populations that includes approximately two-thirds of the Exome Sequencing Project data. In general, an allele frequency in a control population that is greater than expected for the disorder (Table 6) is considered strong support for a benign interpretation for a rare Mendelian disorder (BS1) or, if over 5%, it is considered as stand-alone support (BA1). Furthermore, if the disease under investigation is fully penetrant at an early age and the variant is observed in a well-documented healthy adult individual for a recessive ( homozygous), dominant (heterozygous), or X-linked ( hemizygous) condition, then this is considered strong evidence for a benign interpretation (BS2). If the variant is absent, one should confirm that the read depth in the database is sufficient for an accurate call at the variant site. If a variant is absent from (or below the expected carrier frequency if recessive) a large general population or a control cohort (>1,000 individuals) and the population is race-matched to the patient harboring the identified variant, then this observation can be considered a moderate piece of evidence for pathogenicity (PM2). Many benign variants are “private” (unique to individuals or families), however, and therefore absence in a race-matched population is not considered sufficient or even strong evidence for pathogenicity.
通过搜索公共人群数据库(如千人基因组数据库,NHLBI外显子测序数据库,EXAC数据库; 表1),并利用已发表文献中相同种族的对照数据进行基因变异频率分析(译者注: 此条款在指南更新时会有修改),通过分析变异基因在对照人群或普通人群中的携带频率,有助于评估该变异的潜在致病性。NHLBI外显子测序数据库来源于白种人和非裔美国人群,根据其数据覆盖量能够识别是否存在基因变异。尽管千人基因组数据库缺乏评估基因变异能力,但它囊括了更多的种族人群,因此其数据具有更广泛代表性的。EXAC数据库近期发布了一组来源于不同人群的6万多个外显子组的等位基因频率数据,包括了大约三分之二的NHLBI外显子测序数据。一般情况下,某一等位基因在对照人群的频率大于疾病预期人群(表6)时,可认为是罕见孟德尔疾病良性变异的强证据(BS1),如果频率超过5%时,则可认为是良性变异的独立证据(BA1)。此外,如果疾病发生在早期,且变异在健康成人中以隐性(纯合子)、显性(杂合子)或X-连锁(半合子)的状态存在,那么这就是良性变异的强证据(BS2)。如果数据库中未能检出变异的存在,应该确认建立该数据库采用的测序读长深度是否足以检测出该位点上的变异。如果在一个大样本的普通人群或队列数据的对照人群(>1000人)中变异不存在(或隐性遗传的突变频率是低频),并且携带此变异的患者与对照人群为同一种族,那么可以认为该变异是致病性的中等证据(PM2)。许多良性变异是“个体化的”(即个人或家系独有的),因此即使在相同种族的人群中缺乏也不能作为致病性的充足甚至强的证据。
The use of population data for case–control comparisons is most useful when the populations are well phenotyped, have large frequency differences, and the Mendelian disease under study is early onset. Patients referred to a clinical laboratory for testing are likely to include individuals sent to “rule out” a disorder, and thus they may not qualify as well-phenotyped cases. When using a general population as a control cohort, the presence of individuals with subclinical disease is always a possibility. In both of these scenarios, however, a case–control comparison will be underpowered with respect to detecting a difference; as such, showing a statistically significant difference can still be assumed to provide supportive evidence for pathogenicity, as noted above. By contrast, the absence of a statistical difference, particularly with extremely rare variants and less penetrant phenotypes, should be interpreted cautiously.
当孟德尔遗传病表型显著、频率差异大且是早期发病时,使用通过“病例-对照”人群研究获得的变异数据库进行变异分析是最有效的。临床实验室检测的患者可能包括“排除”某一疾病的个体,因此他们可能不能作为表型显著的病例; 当使用普通人群作为对照群体时,具有亚临床疾病的个体总是可能存在的。在这两种情况下,认为检测出的变异致病性证据不充分。变异频率有统计学显著差异可以假定为致病性的支持证据。与此相反,对于统计差异不显著,特别是极为罕见变异和不明显的表型,应谨慎解释。
Odds ratios (ORs) or relative risk is a measure of association between a genotype (i.e., the variant is present in the genome) and a phenotype (i.e., affected with the disease/ outcome) and can be used for either Mendelian diseases or complex traits. In this guideline we are addressing only its use in Mendelian disease. While relative risk is different from the OR, relative risk asymptotically approaches ORs for small probabilities. An OR of 1.0 means that the variant does not affect the odds of having the disease, values above 1.0 mean there is an association between the variant and the risk of disease, and those below 1.0 mean there is a negative association between the variant and the risk of disease. In general, variants with a modest Mendelian effect size will have an OR of 3 or greater, whereas highly penetrant variants will have very high ORs; for example, APOE E4/E4 homozygotes compared with E3/E3 homozygotes have an OR of 13 (https://www.tgen. org/home/education-outreach/past-summer-interns/2012- summer-interns/erika-kollitz.aspx#.VOSi3C7G_vY). However, the confidence interval (CI) around the OR is as important as the measure of association itself. If the CI includes 1.0 (e.g., OR = 2.5, CI = 0.9–7.4), there is little confidence in the assertion of association. In the above APOE example the CI was ~10–16. Very simple OR calculators are available on the Internet (e.g., http://www.hutchon.net/ConfidOR.htm/ and http://easycalculation.com/statistics/odds-ratio.php/).
比值比(OR)或相对风险用于衡量基因型(即存在于基因组中的变异)和表型(即所患疾病/结果)之间的关联,适用于任何孟德尔疾病或复杂疾病。本指南只涉及其在孟德尔疾病中的使用。相对风险与OR不同,但概率较小时相对风险近似等于OR。OR值为1.0意味着该变异与疾病风险不相关,大于1.0意味着变异与疾病风险正相关,小于1.0意味着变异与疾病风险负相关。一般情况下,具有孟德尔中等效应的变异,其OR值为3或者更大,高度外显的变异具有非常高的OR值,例如,APOE基因E4/E4纯合子与E3/E3纯合子相比,OR值为13(https://www.tgen.org/home/education-outreach/past-summer-interns/2012-summer-interns/erika-kollitz.aspx#.VOSi3C7G_vY)。OR值的置信区间(confidence interval,CI)也是一个重要的衡量工具。如果CI中包括1.0(如OR=2.5,CI=0.9~7.4),则关联的可信度很小。在上面APOE的例子中,CI为10~16。在线可获得简单的OR值计算器(http://www.hutchon.net/ConfidOR.htm/and http://easycalculation.com/statistics/odds-ratio.php/)。
4.6 PM1 热点突变和/或关键的、得到确认的功能域
Certain protein domains are known to be critical to protein function, and all missense variants in these domains identified to date have been shown to be pathogenic. These domains must also lack benign variants. In addition, mutational hotspots in less well-characterized regions of genes are reported, in which pathogenic variants in one or several nearby residues have been observed with greater frequency. Either evidence can be considered moderate evidence of pathogenicity.
某些蛋白结构域对蛋白质的功能起到了关键作用,如果在这些结构域上发现的所有错义突变均已被证实为致病突变,且这些结构域中一定没有已知的良性突变,那么这就能作为致病的中等证据。此外,基因中某些功能尚未确定的区域已被证实存在许多突变热点,若突变发生在基因突变热点上,且一个或多个邻近残基中存在较高频率的已知致病突变,那么这也能作为致病的中等证据。
4.7 PM3 BP2 顺式/反式检测
Testing parental samples to determine whether the variant occurs in cis (the same copy of the gene) or in trans (different copies of the gene) can be important for assessing pathogenicity. For example, when two heterozygous variants are identified in a gene for a recessive disorder, if one variant is known to be pathogenic, then determining that the other variant is in trans can be considered moderate evidence for pathogenicity of the latter variant (PM3). In addition, this evidence could be upgraded to strong if there are multiple observations of the variant in trans with other pathogenic variants. If the variant is present among the general population, however, a statistical approach would be needed to control for random co-occurrence. By contrast, finding the second variant in cis would be supporting, though not definitive, evidence for a benign role (BP2). In the case of uncertain pathogenicity of two heterozygous variants identified in a recessive gene, then the determination of the cis versus trans nature of the variants does not necessarily provide additional information with regard to the pathogenicity of either variant. However, the likelihood that both copies of the gene are impacted is reduced if the variants are found in cis.
检测双亲样本以确定变异在基因上以顺式(in cis)(位于基因的同一拷贝)或是反式(in trans)(位于基因的不同拷贝)方式排列,这对评估变异的致病性非常重要。例如,当两个杂合变异发生在隐性遗传病的致病基因上时,如果已知其中一个变异为致病变异,那么当另一个待分类变异与其呈反式排列时,这可以作为待分类变异的中等致病证据(PM3)。另外,若待分类变异与多个已知致病变异均呈反式排列,则该证据可升级为强致病证据。但是,若待分类变异在普通人群中存在,则需要用统计学方法判断该现象是否为随机共发生事件。相反,当已知致病变异与另一个待分类变异呈顺式排列时,这可以作为待分类变异的良性支持证据(BP2)。如果发生在隐性遗传病致病基因上的两个杂合变异的致病性均未知,那么确定它们以顺式或是反式排列,并不能为判断其中任一变异的致病性提供更多信息。但是,如果两者以顺式排列,则该基因两个拷贝均受影响的可能性将会降低。
In the context of dominant disorders the detection of a variant in trans with a pathogenic variant can be considered supporting evidence for a benign impact (BP2) or, in certain well-developed disease models, may even be considered standalone evidence, as has been validated for use in assessing CFTR variants.
对于显性遗传病而言,若待分类变异与致病变异呈反式排列,则可作为该变异的良性支持证据(BP2); 对于特定研究成熟的疾病模型,甚至可以考虑将其作为独立良性证据(如CFTR相关变异的评估)。
4.8 PM4 BP3 由于框内缺失/插入和终止密码子丧失导致的蛋白长度改变
The deletion or insertion of one or more amino acids as well as the extension of a protein by changing the stop codon to an amino acid codon (e.g., a stop loss variant) is more likely to disrupt protein function compared with a missense change alone as a result of length changes in the protein. Therefore, in-frame deletions/insertions and stop losses are considered moderate evidence of pathogenicity. The larger the deletion, insertion, or extension, and the more conserved the amino acids are in a deleted region, the more substantial is the evidence to support pathogenicity. By contrast, small in-frame deletions/insertions in repetitive regions, or regions that are not well conserved in evolution, are less likely to be pathogenic.
相较于单一的错义突变所导致的蛋白质长度变化,一个或多个氨基酸的缺失或插入、以及由终止密码子变为翻译氨基酸的密码子(如终止密码子丢失)而导致的蛋白质延长更可能破坏蛋白质功能。因此,框内缺失/插入以及终止密码子丢失可作为中等致病证据。缺失、插入或延伸范围越大,缺失区域的氨基酸越保守,则支持致病的证据越强。相反,在重复区域或在进化中不是很保守的区域中小的框内缺失/插入是致病的可能性较小。
4.9 PM5 同一位置新的错义变异
A novel missense amino acid change occurring at the same position as another pathogenic missense change (e.g., Trp38Ser and Trp38Leu) is considered moderate evidence but cannot be assumed to be pathogenic. This is especially true if the novel change is more conservative compared with the established pathogenic missense variant. Also, the different amino acid change could lead to a different phenotype. For example, different substitutions of the Lys650 residue of the FGFR3 gene are associated with a wide range of clinical phenotypes: p.Lys650Gln or p.Lys650Asn causes mild hypochondroplasia; p.Lys650Met causes severe achondroplasia with developmental delay and acanthosis nigricans; and thanatophoric dysplasia type 2, a lethal skeletal dysplasia, arises from p.Lys650Glu.
如果一个新发错义突变发生在一已知致病突变导致相同氨基酸改变的位置上(如Trp38Ser和Trp38Leu),那么可作为中等致病证据(但不能假定一定是致病的),尤其当新的突变比已知致病错义突变更保守时。此外,不同的氨基酸变化可能导致不同的表型。例如,FGFR3基因编码的Lys650残基的不同变化与不同的临床表型相关: p.Lys650Gln或p.Lys650Asn会导致轻度软骨发育不良; p.Lys650Met会导致严重的软骨发育不全伴发育迟缓和黑棘皮病; p.Lys650Glu会导致2型发育异常及致命的骨骼发育不良。
4.10 PP1 BS4 共分离分析
Care must be taken when using segregation of a variant in a family as evidence for pathogenicity. In fact, segregation of a particular variant with a phenotype in a family is evidence for linkage of the locus to the disorder but not evidence of the pathogenicity of the variant itself. A statistical approach has been published with the caveat that the identified variant may be in linkage disequilibrium with the true pathogenic variant in that family. Statistical modeling takes into account age-related penetrance and phenocopy rates, with advanced methods also incorporating in silico predictions and co-occurrence with a known pathogenic variant into a single quantitative measure of pathogenicity. Distant relatives are important to include because they are less likely to have both the disease and the variant by chance than members within a nuclear family. Full gene sequencing (including entire introns and 5′ and 3′ untranslated regions) may provide greater evidence that another variant is not involved or identify additional variants to consider as possibly causative. Unless the genetic locus is evaluated carefully, one risks misclassifying a nonpathogenic variant as pathogenic.
在使用家系中变异的共分离现象作为致病性证据时需谨慎。事实上,一个与某种表型相关的特定变异在某一家系中的共分离现象是位点与疾病连锁的证据,而不是变异本身致病性的证据。一个已经发表的统计方法显示,在某个家系中鉴定的变异可能与真正的致病变异是连锁不平衡的。统计模型考虑到了年龄相关的外显率和拟表型率,一些新的方法也将生物信息分析预测以及与已知致病突变共存作为致病性的单独定量指标。将远亲纳入统计之中是很重要的,因为与核心家系成员相比,他们不太可能同时有该疾病和变异。对整个基因进行测序(包括整个内含子和5′和3′非编码区)可排查其他致病变异或另一个可能致病的变异的存在。除非仔细评估基因位点,否则非致病变异可能被错误地认为是致病变异。
When a specific variant in the target gene segregates with a phenotype or disease in multiple affected family members and multiple families from diverse ethnic backgrounds, linkage disequilibrium and ascertainment bias are less likely to confound the evidence for pathogenicity. In this case, this criterion may be taken as moderate or strong evidence, depending on the extent of segregation, rather than supporting evidence.
当目标基因的特定变异在多个患病的家系成员中以及不同种族背景的多个家系中与表型或疾病共分离时,则其作为致病的证据不太会受到连锁不平衡和确认偏倚的影响。在这种情况下,该标准可以作为中等或强致病证据而不是支持性证据,其强度取决于共分离的程度。
On the other hand, lack of segregation of a variant with a phenotype provides strong evidence against pathogenicity. Careful clinical evaluation is needed to rule out mild symptoms of reportedly unaffected individuals, as well as possible phenocopies (affected individuals with disease due to a nongenetic or different genetic cause). Also, biological family relationships need to be confirmed to rule out adoption, nonpaternity, sperm and egg donation, and other nonbiological relationships. Decreased and age-dependent penetrance also must be considered to ensure that asymptomatic family members are truly unaffected.
另一方面,一个变异与表型并不共分离时,为其非致病的强证据。需要进行仔细的临床评估来排除正常个体的轻度症状和可能的拟表型(患者表型由非遗传或不同的遗传原因引起)。此外,需确认生物学家庭关系来排除收养、非生父、精子和卵子捐献以及其他非生物学关系。同时,外显率下降和年龄依赖性的外显率也必须考虑,以确保无症状家系成员是真正的无症状。
Statistical evaluation of cosegregation may be difficult in the clinical laboratory setting. If appropriate families are identified, clinical laboratories are encouraged to work with experts in statistical or population genetics to ensure proper modeling and to avoid incorrect conclusions of the relevance of the variant to the disease.
在临床实验室进行共分离的统计评估可能并不容易,当鉴定了合适的家系时,为了确保建模合适,并避免得出变异与疾病相关性的错误结论,鼓励临床实验室与统计或群体遗传学专家合作。
4.11 PP2 BP1 变异谱
Many genes have a defined spectrum of pathogenic and benign variation. For genes in which missense variation is a common cause of disease and there is very little benign variation in the gene, a novel missense variant can be considered supporting evidence for pathogenicity (PP2). By contrast, for genes in which truncating variants are the only known mechanism of variant pathogenicity, missense variants can be considered supporting evidence for a benign impact (BP1). For example, truncating variants in ASPM are the primary type of pathogenic variant in this gene, which causes autosomal recessive primary microcephaly, and the gene has a high rate of missense polymorphic variants. Therefore missense variants in ASPM can be considered to have this line of supporting evidence for a benign impact.
许多基因具有明确的致病变异和良性变异谱。在某些基因中,错义突变是导致疾病的常见原因,且该基因上的良性突变非常少,那么这种基因上的新发错义突变可作为致病变异的支持证据(PP2)。相反,有些基因致病的唯一已知变异是截短突变,该基因上的新发错义突变可作为良性的支持证据(BP1)。例如,ASPM基因的截短变异是该基因引起常染色体隐性遗传小头畸形的主要致病变异类型,且该基因发生错义多态性突变的频率高,因此ASPM基因上的错义变异可认为是良性影响的支持证据。
4.12 PP3 BP4 生物信息分析数据
Not overestimating computational evidence is important, particularly given that different algorithms may rely on the same (or similar) data to support predictions and most algorithms have not been validated against well-established pathogenic variants. In addition, algorithms can have vastly different predictive capabilities for different genes. If all of the in silico programs tested agree on the prediction, then this evidence can be counted as supporting. If in silico predictions disagree, however, then this evidence should not be used in classifying a variant. The variant amino acid change being present in multiple nonhuman mammalian species in an otherwise well-conserved region, suggesting the amino acid change would not compromise function, can be considered strong evidence for a benign interpretation. One must, however, be cautious about assuming a benign impact in a nonconserved region if the gene has recently evolved in humans (e.g., genes involved in immune function).
不能过分相信生物信息分析所得到的结果,特别是不同的生物信息算法依赖于相同或相近的数据进行预测,并且大多数生物信息算法未被已知致病变异验证过。此外,相同算法对不同的基因的预测结果可能完全不同。如果不同种类算法的分析预测结果一致,那么生物信息分析结果可以作为支持的证据。如果绝大多数算法的预测结果不一致,则这些预测的结果不能用于对变异进行分类。若某一变异引起的氨基酸改变,在多个非人哺乳动物物种不太保守的区域中出现,说明该变异可能不会损害功能,可以作为良性解读的强的证据。然而,如果某基因已在人类中发生进化(如参与免疫功能的基因),那么在判定该基因在非保守区域中发生的变异为良性时必须小心。
4.13 PP4 表型支持
In general, the fact that a patient has a phenotype that matches the known spectrum of clinical features for a gene is not considered evidence for pathogenicity given that nearly all patients undergoing disease-targeted tests have the phenotype in question. If the following criteria are met, however, the patient’s phenotype can be considered supporting evidence: (i) the clinical sensitivity of testing is high, with most patients testing positive for a pathogenic variant in that gene; (ii) the patient has a welldefined syndrome with little overlap with other clinical presentations (e.g., Gorlin syndrome including basal cell carcinoma, palmoplantar pits, odontogenic keratocysts); (iii) the gene is not subject to substantial benign variation, which can be determined through large general population cohorts (e.g., Exome Sequencing Project); and (iv) family history is consistent with the mode of inheritance of the disorder.
考虑到几乎所有接受疾病针对性测试的患者都有某种表型,通常,不将患者表型与某个基因临床特征谱匹配作为判断致病的证据。但是,如果满足以下条件,患者的表型可作为支持证据: (i) 临床检测的灵敏度高,大多数带有该基因致病突变的患者都被检测为阳性; (ii) 患者有某种明确的综合症的症状,与其他临床表现几乎无重叠(如戈尔林综合征包括基底细胞癌、掌跖坑和牙源性角化); (iii) 该基因通常不存在太多的良性变异(可通过外显子组等人群测序确定的良性变异); (iv) 家族史与疾病遗传方式一致。
4.14 PP5 BP6 可靠的来源
There are increasing examples where pathogenicity classifications from a reputable source (e.g., a clinical laboratory with long-standing expertise in the disease area) have been shared in databases, yet the evidence that formed the basis for classification was not provided and may not be easily obtainable. In this case, the classification, if recently submitted, can be used as a single piece of supporting evidence. However, laboratories are encouraged to share the basis for classification as well as communicate with submitters to enable the underlying evidence to be evaluated and built upon. If the evidence is available, this criterion should not be used; instead, the criteria relevant to the evidence should be used.
现在有越来越多可靠来源(如长期专注于某一疾病领域的临床实验室)的致病性分类信息被分享在数据库中,但分类判断所依据的证据往往并未提供或者很难获取。在这种情况下,如果分类信息是近期提交的,那它就可以作为一个单独的支持证据。然而,还是鼓励实验室共享分类的判断依据,并与提交者进行沟通以评估和创建分类证据。如果能获得证据,则不应使用这一条款,而是应该使用相关的证据。
4.15 BP5 对共发变异的观察
When a variant is observed in a case with a clear alternate genetic cause of disease, this is generally considered supporting evidence to classify the variant as benign. However, there are exceptions. An individual can be a carrier of an unrelated pathogenic variant for a recessive disorder; therefore, this evidence is much stronger support for a likely benign variant classification in a gene for a dominant disorder compared with a gene for a recessive disorder. In addition, there are disorders in which having multiple variants can contribute to more severe disease. For example, two variants, one pathogenic and one novel, are identified in a patient with a severe presentation of a dominant disease. A parent also has mild disease. In this case, one must consider the possibility that the novel variant could also be pathogenic and contributing to the increased severity of disease in the proband. In this clinical scenario, observing the novel variant as the second variant would not support a benign classification of the novel variant (though it is also not considered support for a pathogenic classification without further evidence). Finally, there are certain diseases in which multigenic inheritance is known to occur, such as Bardet-Beidel syndrome, in which case the additional variant in the second locus may also be pathogenic but should be reported with caution.
一般情况下,当某一变异是在一个有明确的遗传病因的疾病患者中被观察到时,可作为将该变异解读为良性的证据。不过,也有例外。某一个体可以是某一不相关隐性遗传疾病致病变异的携带者,因此本证据与隐性遗传性疾病相比,更支持显性遗传性疾病基因良性变异的分类。此外,有些疾病当具有多个变异可以导致更严重的疾病。例如,在一个具有严重表型的显性遗传患者中鉴定了两个变异,一个是致病的,一个是新的变异,父母中的一个也有轻微的疾病,这种情况下,必须考虑新的变异致病的可能性,且新的变异使先证者表型加重。在这种临床情况下,观察到的第二个新的变异不应分类为良性变异,(尽管在无进一步证据的前提下也不认为该变异是致病的)。最后,有些疾病已知为多基因遗传模式,如Bardet-Beidel综合征,在第二个基因座位上的额外变异也有可能是致病的,但应谨慎进行报告。
4.16 BP7 同义变异
There is increasing recognition that splicing defects, beyond disruption of the splice consensus sequence, can be an important mechanism of pathogenicity, particularly for genes in which loss of function is a common mechanism of disease. Therefore, one should be cautious in assuming that a synonymous nucleotide change will have no effect. However, if the nucleotide position is not conserved over evolution and splicing assessment algorithms predict neither an impact to a splice consensus sequence nor the creation of a new alternate splice consensus sequence, then a splicing impact is less likely. Therefore, if supported by computational evidence (BP4), one can classify novel synonymous variants as likely benign. However, if computational evidence suggests a possible impact on splicing or there is raised suspicion for an impact (e.g., the variant occurs in trans with a known pathogenic variant in a gene for a recessive disorder), then the variant should be classified as uncertain significance until a functional evaluation can provide a more definitive assessment of impact or other evidence is provided to rule out a pathogenic role.
人们逐渐认识到经典的剪接序列以外的剪接错误是一类重要的致病机制,特别是对那些功能丧失为其常见致病机制的基因。因此,在假设同义核苷酸改变没有影响时应持谨慎态度。然而如果核苷酸位置进化不保守,且剪接评估算法预测其对剪接一致序列没有影响,也不会产生新的经典剪接序列,那么剪接影响的可能性就比较小。因此,如果生物信息分析证据支持(BP4),可将新发同义变异分类为可能良性。然而,如果生物信息分析证据表明剪接可能有影响或怀疑有影响(例如,发生在隐性遗传病致病基因上,且与已知致病突变呈反式排列的变异),那么在有功能评估可以提供更确切的对影响的评估,或者得到其他可排除该变异致病作用的证据之前,该类变异应该归类为意义不明确。
5. 序列变异报导
Writing succinct yet informative clinical reports can be a challenge as the complexity of the content grows from reporting variants in single genes to multigene panels to exomes and genomes. Several guidance documents have been developed for reporting, including full sample reports of the ACMG clinical laboratory standards for next-generation sequencing guidance. Clinical reports are the final product of laboratory testing and often are integrated into a patient’s electronic health record. Therefore, effective reports are concise, yet easy to understand. Reports should be written in clear language that avoids medical genetics jargon or defines such terms when used. The report should contain all of the essential elements of the test performed, including structured results, an interpretation, references, methodology, and appropriate disclaimers. These essential elements of the report also are emphasized by Clinical Laboratory Improvement Amendments regulations and the College of American Pathologists laboratory standards for next-generation sequencing clinical tests.
编写简明而内容丰富的临床报告不是一件容易的事情,因为从检测单个基因,到多基因包,再到外显子组和基因组,变异情况的报告内容复杂程度会大大增加。为规范报告内容已出台了一些指南文件,包括符合ACMG临床实验室标准的新一代测序检测完整报告示例。临床报告是实验室检测结果的最终体现,通常会放入到患者的电子健康档案中。因此,有效的报告应该是简明扼要且易于理解的。报告应该使用清晰的语言书写,避免使用医学遗传学术语,当必须要使用时需指明所用术语的定义。报告应包含所有的检测基本要素,包括结构化的结果、解释、参考文献、检测方法和适当的免责声明。《临床实验室改进法案》(CLIA)以及美国病理学家学会在针对新一代测序临床实验标准中,也强调了上述基本要素。
5.1 结果
The results section should list variants using HGVS nomenclature (see Nomenclature). Given the increasing number of variants found in genetic tests, presenting the variants in tabular form with essential components may best convey the information. These components include nomenclature at both the nucleotide (genomic and complementary DNA) and protein level, gene name, disease, inheritance, exon, zygosity, and variant classification. An example of a table to report structured elements of a variant is found in the Supplementary Appendix S1 online. Parental origin may also be included if known. In addition, if specific variants are analyzed in a genotyping test, the laboratory should specifically note the variants interrogated, with their full description and historical nomenclature if it exists. Furthermore, when reporting results from exome or genome sequencing, or occasionally very large disease-targeted panels, grouping variants into categories such as “Variants in Disease Genes with an Established Association with the Reported Phenotype,” “Variants in Disease Genes with a Likely Association with the Reported Phenotype,” and (where appropriate) “Incidental (Secondary) Findings” may be beneficial.
结果部分应根据HGVS命名规则(见命名部分)列出变异。考虑到在基因检测中发现的变异数目越来越多,以包含基本内容的表格呈现变异结果可能是传达信息的最好方法。这些基本内容包括在核苷酸(基因组和cDNA)和蛋白质水平的命名、基因名称、疾病、遗传模式、外显子、合子性及变异的分类. 若亲本来源明确,也可包括在内。此外,如果变异是通过基因分型检测的,实验室应特别注明受检变异的完整描述及曾用名。当报告外显子组或全基因组测序结果,或偶尔报告包含基因数目较多的疾病基因包检测结果时,将变异按“与表型明确相关的疾病基因的变异”、“与表型可能相关的疾病基因的变异”及(在适当情况下)“附带(次要)发现”进行分类可能有益。
5.2 解读
The interpretation should contain the evidence supporting the variant classification, including its predicted effect on the resultant protein and whether any variants identified are likely to fully or partially explain the patient’s indication for testing. The report also should include any recommendations to the clinician for supplemental clinical testing, such as enzymatic/ functional testing of the patient’s cells and variant testing of family members, to further inform variant interpretation. The interpretation section should address all variants described in the results section but may contain additional information. It should be noted whether the variant has been reported previously in the literature or in disease or control databases. The references, if any, that contributed to the classification should be cited where discussed and listed at the end of the report. The additional information described in the interpretation section may include a summarized conclusion of the results of in silico analyses and evolutionary conservation analyses. However, individual computational predictions (e.g., scores, terms such as “damaging”) should be avoided given the high likelihood of misinterpretation by health-care providers who may be unfamiliar with the limitations of predictive algorithms (see In Silico Predictive Programs, above). A discussion of decreased penetrance and variable expressivity of the disorder, if relevant, should be included in the final report. Examples of how to describe evidence for variant classification on clinical reports are found in the Supplementary Appendix S1 online.
解读应包含对变异检测结果进行分类的证据,包括编码蛋白的功能影响预测,以及检测所发现的变异是否可能全部或部分地解释患者的临床表型。报告也应包括对临床医生的建议,这些建议包括一些需补充的临床检测,如对患者进行细胞酶学/功能的检测,以及对患者家系其他成员进行的变异检测,以便为进一步解读变异检测结果提供支持。解读应当包括检测结果部分描述的全部变异,以及其他附加信息。对于各个变异需要注明是否已经在先前的文献、疾病病例或对照数据库中有过报道。在报告结尾处需要列出对变异检测结果分类时所引用的全部参考文献和信息。解读部分其他的附加信息可以包括对变异位点进行进化保守性分析的结果总结。由于医务工作者可能不熟悉预测算法的局限性(详见上文“3.4生物信息学计算预测程序”小节),因此,应该避免报告对个体进行生物信息学预测的计算结果(如分数,诸如“破坏性”之类的术语),以免造成医务工作者对报告产生误解。 如果存在疾病的外显率下降和表现度差异,也需要将有关的讨论包含在最终的报告中。
5.3 方法学
The methods and types of variants detected by the assay and those refractory to detection should be provided in the report. Limitations of the assay used to detect the variants also should be reported. Methods should include those used to obtain nucleic acids (e.g., polymerase chain reaction, capture, wholegenome amplification), as well as those to analyze the nucleic acids (e.g., bidirectional Sanger sequencing, next-generation sequencing, chromosomal microarray, genotyping technologies), because this may provide the health-care provider with the necessary information to decide whether additional testing is required to follow up on the results. The methodology section should also give the official gene names approved by the Human Genome Organization Gene Nomenclature Committee, RefSeq accession numbers for transcripts, and genome build, including versions. For large panels, gene-level information may be posted and referenced by URL. The laboratory may choose to add a disclaimer that addresses general pitfalls in laboratory testing, such as sample quality and sample mix-up.
报告中应说明使用的实验方法、检测所涉及的变异类型、检测过程的难点,以及检测变异所使用的方法的局限性。需要说明的实验方法应包括核酸的获取方法(如聚合酶链式反应、捕获、全基因组扩增等)以及核酸的检测方法(如双向Sanger测序、下一代测序、染色体基因芯片、基因分型技术等),这些信息可以为医务工作者提供必要的信息,以帮助其决定是否需要追加实验来跟进这些检测结果。方法部分还应包括人类基因组组织基因命名委员会批准的正式基因名称、转录本的RefSeq登录号和所参考的基因组版本。对于大的基因包,基因水平的信息可以通过引用URL来加以说明。实验室还可以选择增加对检测过程中常见问题(如样本质量问题、样品混合污染等)的免责声明。
5.4 患者维权团体、临床实验和研究的获取
Although specific clinical guidance for a patient is not recommended for laboratory reports, provision of general information for categories of results (e.g., all positives) is appropriate and helpful. A large number of patient advocacy groups and clinical trials are now available for support and treatment of many diseases. Laboratories may choose to add this information to the body of the report or attach the information so it is sent to the health-care provider along with the report. Laboratories may make an effort to connect the health-care provider to research groups working on specific diseases when a variant’s effect is classified as “uncertain,” as long as Health Insurance Portability and Accountability Act patient privacy requirements are followed.
尽管不提倡在实验室报告中对患者提供具体临床指导,但是在报告中提供对于检测结果分类的总体信息(如全部阳性检测结果)是恰当且有益的。大量病人群体和临床试验现在可用于多种疾病的支持和治疗。实验室可以选择将此信息添加到报告的正文或附加信息,并且与报告一起发送给医务工作者。在遵守医疗保险便携性和责任法案(HIPAA)保护患者隐私的前提下,当某一变异检测结果被归为意义不明确时,实验室可尝试帮助医务工作者和特定的疾病研究小组建立联系。
5.5 变异再分析
As evidence on variants evolves, previous classifications may later require modification. For example, the availability of variant frequency data among large populations has led many uncertain significance variants to be reclassified as benign, and testing additional family members may result in the reclassification of variants.
随着新的变异证据增加,现有的分类标准需要修订。例如,当大样本的有效的人群变异频率被报道后,许多原本意义不明确的变异,可以因为明确意义而进行重新分类,而检测家系中其他成员的结果也可以导致重新分类。
As the content of sequencing tests expands and the number of variants identified grows, expanding to thousands and millions of variants from exome and genome sequencing, the ability for laboratories to update reports as variant knowledge changes will be untenable without appropriate mechanisms and resources to sustain those updates. To set appropriate expectations with health-care providers and patients, laboratories should provide clear policies on the reanalysis of data from genetic testing and whether additional charges for reanalysis may apply. Laboratories are encouraged to explore innovative approaches to give patients and providers more efficient access to updated information.
随着检测变异数量的增加及检测范围的扩大,无论是全外显子检测还是全基因组测序,都可以得到数以百万的变异信息量。如果实验室缺乏有效的分析方法和足够的文献数据库支撑,将无法进行变异再分析。为了满足医务人员和患者的实际需求,实验室应该开展基因检测数据再分析,并明确再分析是否产生额外费用。应该鼓励实验室为帮助医务人员和患者而不断开发更新信息的新途径。
For reports containing variants of uncertain significance in genes related to the primary indication, and in the absence of updates that may be proactively provided by the laboratory, it is recommended that laboratories suggest periodic inquiry by health-care providers to determine whether knowledge of any variants of uncertain significance, including variants reported as likely pathogenic, has changed. By contrast, laboratories are encouraged to consider proactive amendment of cases when a variant reported with a near-definitive classification (pathogenic or benign) must be reclassified. Regarding physician responsibility, see the ACMG guidelines on the duty to recontact.
当报告中有针对主要指征的基因中存在临床意义不明的变异,在实验室又无法及时提供更新的数据时,建议医务人员定期查询其不明意义的变异结果是否被更改。另一方面,鼓励实验室在对变异的分类有重要变化时(如致病性或良性的变异被修改)必须主动及时地更新报告。关于医生对病人报告更新方面的责任,可详见ACMG有关指南。
5.6 变异的验证
Recommendations for the confirmation of reported variants is addressed elsewhere. Except as noted, confirmation studies using an orthogonal method are recommended for all sequence variants that are considered to be pathogenic or likely pathogenic for a Mendelian disorder. These methods may include, but are not limited to, re-extraction of the sample and testing, testing of parents, restriction enzyme digestion, sequencing the area of interest a second time, or using an alternate genotyping technology.
关于变异验证的建议已在其他地方阐述了。再次重申,对于孟德尔疾病的致病或可能致病变异需进行正交法验证。具体方法包括但不限于以下几种: 重新取样和检测、检测父母的变异情况、限制性内切酶消化、对于目标区域重新测序或使用另一种基因分型技术。
6. 特殊考虑
6.1 对临床意义不明确的基因(GUS)中的变异的评估和报告
Genome and exome sequencing are identifying new genotype– phenotype connections. When the laboratory finds a variant in a gene without a validated association to the patient’s phenotype, it is a GUS. This can occur when a gene has never been associated with any patient phenotype or when the gene has been associated with a different phenotype from that under consideration. Special care must be taken when applying the recommended guidelines to a GUS. In such situations, utilizing variant classification rules developed for recognized genotype– phenotype associations is not appropriate. For example, when looking across the exome or genome, a de novo observation is no longer strong evidence for pathogenicity given that all individuals are expected to have approximately one de novo variant in their exome or 100 in their genome. Likewise, thousands of variants across a genome could segregate with a significant logarithm of the odds (LOD) score. Furthermore, many deleterious variants that are clearly disruptive to a gene or its resultant protein (nonsense, frameshift, canonical ±1,2 splice site, exonlevel deletion) may be detected; however, this is insufficient evidence for a causative role in any given disease presentation.
基因组和外显子组测序正在不断鉴定出新的基因型-表型关联。当实验室发现某个基因的变异,但尚未证实此基因与病人的表型有关联,该变异称为GUS变异。这种情况可出现在当一个基因从未与任何病人表型相关联时,或者与此基因相关联的表型不同于正在被考虑的表型。当推荐的指南应用于GUS时必须特别注意。在这种情况下,由于目前指南中的变异分类规则适用于已经明确的基因型-表型关联,但并不适合未知的情况。例如,纵观外显子组或基因组,考虑到所有个体的外显子组中预计约有1个新发变异或基因组中约有100个新发变异,新发变异的发现不再是致病性的强有力证据。同样地,整个基因组中成千上万个变异可与显著的LOD值共分离。此外,许多明显破坏基因或其蛋白的有害变异(无义、移码、典型±1,2剪接位点、外显子水平缺失)可能被检测出来,然而,在对任何疾病表型的解释中,这些变异都不是充分的致病证据。
Variants found in a GUS may be considered as candidates and reported as “variants in a gene of uncertain significance.” These variants, if reported, should always be classified as uncertain significance. Additional evidence would be required to support the gene’s association to disease before any variant in the gene itself can be considered pathogenic for that disease. For example, additional cases with matching rare phenotypes and deleterious variants in the same gene would enable the individual variants to be classified according to the recommendations presented here.
GUS中发现的变异可作为候选,并可报告为“意义不明确的基因变异”。如果报道这些变异,应该一直被分类为意义不明确。在任何基因变异可被考虑为疾病的致病原因之前,都需要附加的证据支持基因与疾病的关联。例如,与罕见表型匹配和存在相同基因上存在有害变异的其他病例使得可以根据此指南对某一变异进行分类。
6.2 在健康个体中评估变异或作为偶然发现
Caution must be exercised when using these guidelines to evaluate variants in healthy or asymptomatic individuals or to interpret incidental findings unrelated to the primary indication for testing. In these cases the likelihood of any identified variant being pathogenic may be far less than when performing disease-targeted testing. As such, the required evidence to call a variant pathogenic should be higher, and extra caution should be exercised. In addition, the predicted penetrance of pathogenic variants found in the absence of a phenotype or family history may be far less than predicted based on historical data from patients ascertained as having disease.
当评估在健康或无症状个体中检测到的变异或者解释与主要检测指征无关的偶然发现的变异时,必须谨慎使用此指南。在这些情况下,所识别变异为致病变异的概率可能远低于疾病靶向性检测。正因为如此,当判定这些变异为致病变异时,不仅需要更强的证据支持,而且需要额外谨慎。此外,和基于确诊患者预测的外显率相比,在无相关表型或家族史个体中发现的致病变异的预测外显率可能要低很多。
6.3 线粒体变异
The interpretation of mitochondrial variants other than well-established pathogenic variants is complex and remains challenging; several special considerations are addressed here.
除了明确的致病变异,线粒体变异的解读是复杂且依旧充满挑战的,此处提出了一些特殊的考虑。
The nomenclature differs from standard nomenclature for nuclear genes, using gene name and m. numbering (e.g., m.8993T>C) and p. numbering, but not the standard c. numbering (see also Nomenclature). The current accepted reference sequence is the Revised Cambridge Reference Sequence of the Human Mitochondrial DNA: GenBank sequence NC_012920 gi:251831106.
线粒体变异的命名法与核基因的标准命名法不同,使用基因名和m.编号(如m.8993T>C)和p.编号,而不是标准的c.编号(见命名法)。目前公认的参考序列是人类线粒体DNA修订版剑桥参考序列: 基因库序列NC_012920 gi: 251831106(http://www.mitomap.org/MITOMAP/HumanMitoSeq)。
Heteroplasmy or homoplasmy should be reported, along with an estimate of heteroplasmy of the variant if the test has been validated to determine heteroplasmy levels. Heteroplasmy percentages in different tissue types may vary from the sample tested; therefore, low heteroplasmic levels also must be interpreted in the context of the tissue tested, and they may be meaningful only in the affected tissue such as muscle. Over 275 mitochondrial DNA variants relating to disease have been recorded (http://mitomap.org/bin/view.pl/MITOMAP/WebHome). MitoMap is considered the main source of information related to mitochondrial variants as well as haplotypes. Other resources, such as frequency information (http://www.mtdb.igp.uu.se/), secondary structures, sequences, and alignment of mitochondrial transfer RNAs (http://mamittrna.u-strasbg.fr/), mitochondrial haplogroups (http://www.phylotree.org/)and other information (http://www.mtdnacommunity.org/default.aspx), may prove useful in interpreting mitochondrial variants.
如果已通过检测对异质性水平进行确定,应该对异质性或同质性,以及变异异质性的评估进行报道。不同组织类型的异质性百分比因检测样本的不同而有所改变, 因此,低异质性水平也必须结合所检测组织进行解读,且它们可能仅在受累及的组织中才是有意义的,如肌肉组织。超过275个与疾病相关的线粒体DNA变异已被记录(http://mitomap.org/bin/view.pl/MITOMAP/WebHome)。MitoMap是线粒体变异及单倍型相关信息的主要来源。其他资源,如频率信息(http://www.mtdb.igp.uu.se/)、二级结构、序列和线粒体转运RNA的比对(http://mamittrna.u-strasbg.fr/)、线粒体单倍群(http://www.phylotree.org/)和其他信息(http://www.mtdnacommunity.org/default.aspx),可能在解读线粒体变异时是有用的。
Given the difficulty in assessing mitochondrial variants, a separate evidence checklist has not been included. However, any evidence needs to be applied with additional caution. The genes in the mitochondrial genome encode for transfer RNA as well as for protein; therefore, evaluating amino acid changes is relevant only for genes encoding proteins. Similarly, because many mitochondrial variants are missense variants, evidence criteria for truncating variants likely will not be helpful. Because truncating variants do not fit the known variant spectrum in most mitochondrial genes, their significance may be uncertain. Although mitochondrial variants are typically maternally inherited, they can be sporadic, yet de novo variants are difficult to assess because of heteroplasmy that may be below an assay’s detection level or different between tissues. The level of heteroplasmy may contribute to the variable expression and reduced penetrance that occurs within families. Nevertheless, there remains a lack of correlation between the percentage of heteroplasmy and disease severity. Muscle, liver, or urine may be additional specimen types useful for clinical evaluation. Undetected heteroplasmy may also affect outcomes of case, case–control, and familial concordance studies. In addition, functional studies are not readily available, although evaluating muscle morphology may be helpful (i.e., the presence of ragged red fibers). Frequency data and published studies demonstrating causality may often be the only assessable criteria on the checklist. An additional tool for mitochondrial diseases may be haplogroup analysis, but this may not represent a routine method that clinical laboratories have used, and the clinical correlation is not easy to interpret.
鉴于线粒体变异评估的难度,本指南并未包括单独的证据清单。然而,任何证据的应用均需要格外谨慎。线粒体基因组中的基因编码转运RNA和蛋白质,因此,评估氨基酸的变化仅与蛋白质的编码基因有关。同样地,因为很多线粒体变异是错义突变,截短突变的证据标准可能并不适用。由于截短突变并不符合多数线粒体基因的已知变异谱,其意义可能是不确定的。尽管线粒体变异是典型的母系遗传,它们也可以散发。然而由于异质性可能低于试验检测水平或组织间的差异,新发变异是难以评估的。异质性水平可能是家族内表达差异和外显率降低的原因。尽管如此,异质性百分比和疾病严重程度之间仍缺乏相关性。肌肉、肝脏或尿液可以作为附加样本类型用于临床评估。未检测到的异质性也可能影响病例、病例对照和家系一致性研究的结果。此外,没有现成的功能研究方法,尽管评估肌肉形态可能会有所帮助(即破碎红纤维的存在)。频率数据和已发表的证明因果关系的研究往往是检测报告上唯一的评估标准。单倍群分析可以作为线粒体疾病的另一个工具,但可能不是临床实验室已使用的常规方法,而且临床相关性难以解释。
Consideration should be given to testing nuclear genes associated with mitochondrial disorders because variants in nuclear genes could be causative of oxidative disorders or modulating the mitochondrial variants.
因为核基因变异也可能是氧化疾病的致病原因或起着调节线粒体变异的作用,因此应考虑检测与线粒体疾病相关的核基因。
6.4 药物基因组学
Establishing the effects of variants in genes involved with drug metabolism is challenging, in part because a phenotype is only apparent upon exposure to a drug. Still, variants in genes related to drug efficacy and risk for adverse events have been described and are increasingly used in clinical care. Gene summaries and clinically relevant variants can be found in the Pharmacogenomics Knowledge Base (http://www.pharmgkb.org/). Alleles and nomenclature for the cytochrome P450 gene family is available at http://www.cypalleles.ki.se/. Although the interpretation of PGx variants is beyond the scope of this document, we include a discussion of the challenges and distinctions associated with the interpretation and reporting of PGx results.
确认基因变异在药物代谢中的作用具有挑战性,部分原因在于其表型只有在接触药物后才得以显现。不过,临床上现已报告了各种与药物疗效和副作用风险相关的基因变异,且其数量仍然在不断增加。相关基因的汇总及其有临床意义的变异可查询药物基因组学知识库网站(http://www.pharmgkb.org/)。有关细胞色素P450基因家族等位基因及其命名可查询网站http://www.cypalleles.ki.se/。尽管解读药物基因组变异已超出了本文的范围,还是对与解读及报告药物基因组结果相关的挑战和鉴别进行了讨论。
The traditional nomenclature of PGx alleles uses star (*) alleles, which often represent haplotypes, or a combination of variants on the same allele. Traditional nucleotide numbering using outdated reference sequences is still being applied. Converting traditional nomenclature to standardized nomenclature using current reference sequences is an arduous task, but it is necessary for informatics applications with next-generation sequencing.
传统的药物基因组等位基因命名使用星号(*)标记等位基因,通常用于表示单倍型或同一等位上基因变异的组合。依据旧的参考序列的传统核苷酸编号规则仍然在使用。而将传统命名转换为使用新参考序列的标准命名会是一项艰巨的任务,但这对于下一代测序的信息学分析应用是必要的。
Many types of variants have been identified in PGx genes, such as truncating, missense, deletions, duplications (of functional as well as nonfunctional alleles), and gene conversions, resulting in functional, partially functional (decreased or reduced function), and nonfunctional (null) alleles. Interpreting sequence variants often requires determining haplotype from a combination of variants detected. Haplotypes are typically presumed based on population frequencies and known variant associations rather than testing directly for chromosomal phase (molecular haplotyping).
在药物基因组相关基因上已经鉴定了多种变异类型,如截短、错义、缺失、重复(含功能及非功能等位基因)及基因转换,它们可导致等位基因功能性或部分功能性丧失(功能减退或降低)以及无功能性(无效)等位基因。解读序列变异常需依据由各种遗传变异组合成的单倍型信息。单倍型一般根据人群频率和已知变异关联分析信息进行推定,而非通过直接检测染色体片段(分子单倍型)来实现。
In addition, for many PGx genes (particularly variants in genes coding for enzymes), the overall phenotype is derived from a diplotype, which is the combination of variants or haplotypes on both alleles. Because PGx variants do not directly cause disease, using terms related to metabolism (rapid, intermediate, poor); efficacy (resistant, responsive, sensitive); or “risk,” rather than pathogenic, may be more appropriate. Further nomenclature and interpretation guidelines are needed to establish consistency in this field.
此外,对于许多药物基因组基因(特别是酶编码基因变异),其整体表型取决于二倍型,即两个等位基因上的变异或单倍型组合。由于药物基因组变异并不直接导致疾病,使用代谢(快速、中等及减弱)、疗效(耐药、响应及敏感)或“风险”而非“致病”应更为恰当。需要建立起在本领域保持一致性的专业术语和解读指南。
6.5 常见复杂疾病
Unlike Mendelian diseases, the identification of common, complex disease genes, such as those contributing to type 2 diabetes, coronary artery disease, and hypertension, has largely relied on population-based approaches (e.g., genome-wide association studies) rather than family-based studies. Currently, numerous genome-wide association study reports have resulted in the cataloguing of over 1,200 risk alleles for common, complex diseases and traits. Most of these variants are in nongenic regions, however, and additional studies are required to determine whether any of the variants are directly causal through effects on regulatory elements, for example, or are in linkage disequilibrium with causal variants.
与孟德尔疾病以家系为基础的研究不同,常见复杂疾病(如2型糖尿病、冠心病和高血压)相关基因的鉴定,在很大程度上依赖于以人群为基础的方法(如全基因组关联分析)。目前,大量的全基因组关联研究报告已对1200余种常见复杂疾病和性状的风险等位基因进行了编目. 然而,这些变异大多数位于基因之间的区域,尚需要进一步的研究来确定这些变异是否通过影响调控因子而直接导致疾病,或者与致病变异处于连锁不平衡状态。
Common, complex risk alleles typically confer low relative risk and are meager in their predictive power. To date, the utility of common, complex risk allele testing for patient care has been unclear, and models to combine multiple markers into a cumulative risk score often are flawed and are usually no better than traditional risk factors such as family history, demographics, and nongenetic clinical phenotypes. Moreover, in almost all of the common diseases the risk alleles can explain only up to 10% of the variance in the population, even when the disease has high heritability. Given the complexity of issues, this recommendation does not address the interpretation and reporting of complex trait alleles. We recognize, however, that some of these alleles are identified during the course of sequencing Mendelian genes, and therefore guidance on how to report such alleles when found incidentally is needed. The terms “pathogenic” and “likely pathogenic” are not appropriate in this context, even when the association is statistically valid. Until better guidance is developed, an interim solution is to report these variants as “risk alleles” or under a separate “other reportable” category in the diagnostic report. The evidence for the risk, as identified in the case–control/ genome-wide association studies, can be expressed by modifying the terms, such as “established risk allele,” “likely risk allele,” or “uncertain risk allele,” if desired.
常见复杂风险等位基因通常被赋予较低的相对风险,且预测能力薄弱。迄今为止,常见复杂风险等位基因检测对于患者治疗的效用尚不清楚,将多个指标组合起来进行累计风险评估的模型往往是有缺陷的,通常并不优于家族史、人口统计资料和非遗传性临床表型等传统风险因素。另外,在几乎所有的常见疾病中,风险等位基因仅可解释至多10%的群体变异,即使当疾病有高度遗传度时也是如此。考虑到问题的复杂性,本建议并不涉及复杂性状的等位基因的解读和报告。然而我们认识到,在对孟德尔基因进行测序时可以识别这些等位基因中的一部分,因此需要有偶然发现这些等位基因时如何进行报告的指南。这种情况下,术语“致病的”和“可能致病的”并不适用,即使关联在统计学上是有效的。在建立更好的指南之前,临时的解决办法是将这些变异报告为“风险等位基因”,或在诊断报告中设立一个单独的“其他报告”类别。同病例对照/全基因组关联研究鉴定一样,风险证据可以通过修改术语来表达,如“确定的风险等位基因”,“可能的风险等位基因”或“不确定的风险等位基因”。
6.6 体细胞变异
The description of somatic variants, primarily those observed in cancer cells, includes complexities not encountered with constitutional variants, because the allele ratios are highly variable and tumor heterogeneity can cause sampling differences. Interpretation helps select therapy and predicts treatment response or the prognosis of overall survival or tumor progression–free survival, further complicating variant classification. For the interpretation of negative results, understanding the limit of detection of the sequencing assay (at what allele frequency the variant can be detected by the assay) is important and requires specific knowledge of the tumor content of the sample. Variant classification categories are also different, with somatic variants compared with germ-line variants, with terms such as “responsive,” “ resistant,” “driver,” and “passenger” often used. Whether a variant is truly somatic is confirmed by sequence analysis of the patient’s germ-line DNA. A different set of interpretation guidelines is needed for somatic variants, with tumor-specific databases used for reference, in addition to databases used for constitutional findings. To address this, a workgroup has recently been formed by the AMP.
体细胞变异主要见于癌细胞,因为其等位基因比值高度可变,且肿瘤异质性也可导致取样差异。在描述其变异时,具有原发性变异所没有的复杂性。变异的解读有助于选择治疗方案和预测治疗效果、也应用于评估整体生存率或肿瘤无进展生存期,因而体细胞变异的分类更加复杂。在对阴性结果解读时,了解测序分析的检测方法局限性(变异可在何种等位基因频率时被检测到)至关重要,此外也需要了解样本中肿瘤含量的特定信息。与胚系变异相比,体细胞变异的分类类别也不同,通常使用“敏感”、“拮抗”、“驱动”和“伴随”等术语。一个变异是否是体细胞变异需要通过患者胚系DNA的序列分析来证实。体细胞变异还需要另外的解读指南,除了参考原发性突变的数据库以外,还需要肿瘤特异性数据库作为参考。为了解决这个问题,最近AMP已经成立了一个工作组。
7. 医疗工作者如何使用这些指南和建议
The primary purpose of clinical laboratory testing is to support medical decision making. In the clinic, genetic testing is generally used to identify or confirm the cause of disease and to help the health-care provider make individualized treatment decisions including the choice of medication. Given the complexity of genetic testing, results are best realized when the referring health-care provider and the clinical laboratory work collaboratively in the testing process.
临床实验室检测的主要目的是为医疗决策提供依据。在临床上,基因检测一般用于识别或确认疾病的原因,并帮助医务工作者做出个性化的治疗决策,包括用药的选择。鉴于基因检测的复杂性,检测过程中需相关医务工作者和临床实验室协作才能得到最佳结果。
When a health-care provider orders genetic testing, the patient’s clinical information is integral to the laboratory’s analysis. As health-care providers increasingly utilize genomic (exome or genome) sequencing, the need for detailed clinical information to aid in interpretation assumes increasing importance. For example, when a laboratory finds a rare or novel variant in a genomic sequencing sample, the director cannot assume it is relevant to a patient just because it is rare, novel, or de novo. The laboratory must evaluate the variant and the gene in the context of the patient’s and family’s history, physical examinations, and previous laboratory tests to distinguish between variants that cause the patient’s disorder and those that are incidental (secondary) findings or benign. Indeed, accurate and complete clinical information is so essential for the interpretation of genome-level DNA sequence findings that the laboratory can reasonably refuse to proceed with the testing if such information is not provided with the test sample.
当医务工作者提出基因检测需求时,需将患者的临床信息提供给实验室。由于医务工作者越来越多地使用基因组(全外显子组或全基因组)测序,而详细的临床信息有助于对检测结果的解读,因此向实验室提供临床信息就变得越来越重要。例如,当一个实验室在基因组测序样品中发现一个罕见或新发的变异时,实验室负责人不能仅因为该变异是罕见的、新发现的或者新发的来确定它的致病性。该实验室必须通过患者的病史、家族史、体格检查和前期实验室检查对变异和基因进行评估,进而区分致病变异和其他偶然(次要)发现或良性变异。事实上,准确和完整的临床信息对于基因组水平DNA序列检测结果的解读是不可或缺的,若待测样品不能提供此类信息,实验室可以合理拒绝继续进行检测。
For tests that cover a broad range of phenotypes (large panels, exome and genome sequencing) the laboratory may find candidate causative variants. Further follow-up with the health-care provider and patient may uncover additional evidence to support a variant. These additional phenotypes may be subclinical, requiring additional clinical evaluation to detect (e.g., temporal bone abnormalities detected by computed tomography in a hearing-impaired patient with an uncertain variant in SLC26A4, the gene associated with Pendred syndrome). In addition, testing other family members to establish when a variant is de novo, when a variant cosegregates with disease in the family, and when a variant is in trans with a pathogenic variant in the same recessive disease-causing gene is valuable. Filtering out or discounting the vast majority of variants for dominant diseases when they can be observed in healthy relatives is possible, making the interpretation much more efficient and conclusive. To this end, it is strongly recommended that every effort be made to include parental samples along with that of the proband, so-called “trio” testing (mother, father, affected child), in the setting of exome and genome sequencing, particularly for suspected recessive or de novo causes. Obviously this will be easier to achieve for pediatric patients than for affected adults. In the absence of one or both parents, the inclusion of affected and unaffected siblings can be of value.
利用如高通量的靶向测序、全外显子组和全基因组测序等覆盖广泛表型的方法进行检测,实验室可能会发现候选的致病变异。对医务工作者和患者后续的随访可能会发现更多的证据来支持某一变异的致病性。这些补充的表型信息可能是亚临床症状,需要进一步完善相关的临床检测(例如,一个在SLC26A4基因(与Pendred综合征相关的基因)上有不确定变异的听力受损患者,需要进行CT检查判断其有无颞骨异常)。此外,当发现一个变异可能是新发变异,或者当一个变异在家系中与表型共分离,或者在隐性遗传致病基因中一个变异与另一个致病变异处于反式位置时,必须在其他家系成员中进行验证。在显性遗传性疾病的情况下,在健康亲属中观察到的绝大部分变异可以被过滤或删减,这样可使解读更加有效和准确。为此,强烈建议在开展外显子组或基因组测序时,尽力做到“核心家系”检测(即母亲、父亲、患病儿童),尤其是对怀疑有隐性遗传或新发变异的患者。与成人患者相比,这显然在儿科患者中更易实现。在没有父母一方或双方时,纳入患病和正常的兄弟姐妹也是有意义的。
Many genetic variants can result in a range of phenotypic expression (variable expressivity), and the chance of disease developing may not be 100% (reduced penetrance), further underscoring the importance of providing comprehensive clinical data to the clinical laboratory to aid in variant interpretation. Ideally, it is recommended that clinical data be deposited into, and shared via, centralized repositories as allowable by Health Insurance Portability and Accountability Act and institutional review board regulations. Importantly, referring health-care providers can further assist clinical laboratories by recruiting DNA from family members in scenarios where their participation will be required to interpret results, (e.g., when evaluating cosegregation with disease using affected family members, genotyping parents to assess for de novo occurrence and determining the phase of variants in recessive disorders using first-degree relatives).
许多遗传变异会导致一系列表型(不同程度的表现度),疾病发生的机率也可能不是100%(外显率降低),这些均进一步强调了向临床实验室提供全面的临床数据来帮助解读变异的重要性。在理想的情况下,建议应依据医疗保险可携性和责任法案(HIPAA)和机构审查委员会条例,将临床数据存入并通过集中存储库共享。重要的是,当家庭成员的信息对于解读结果是必需的时候,相关医务工作者可以进一步帮助临床实验室收集家庭成员的DNA(例如,当评估家系患者与疾病共分离时,父母的基因型分析可用来评估新发变异的发生,一级亲属可用来确定隐性遗传疾病变异的同线或异线性)。
A key issue for health-care providers is how to use the evidence provided by genetic testing in medical management decisions. Variant analysis is, at present, imperfect, and the variant category reported does not imply 100% certainty. In general, a variant classified as pathogenic using the proposed classification scheme has met criteria informed by empirical data such that a health-care provider can use the molecular testing information in clinical decision making. Efforts should be made to avoid using this as the sole evidence of Mendelian disease; it should be used in conjunction with other clinical information when possible. Typically, a variant classified as likely pathogenic has sufficient evidence that a health-care provider can use the molecular testing information in clinical decision making when combined with other evidence of the disease in question. For example, in the prenatal setting an ultrasound may show a key confirmatory finding; in postnatal cases, other data such as enzyme assays, physical findings, or imaging studies may conclusively support decision making. However, it is recommended that all possible follow-up testing, as described above, be pursued to generate additional evidence related to a likely pathogenic variant because this may permit the variant to be reclassified as pathogenic. A variant of uncertain significance should not be used in clinical decision making. Efforts to resolve the classification of the variant as pathogenic or benign should be undertaken. While this effort to reclassify the variant is underway, additional monitoring of the patient for the disorder in question may be prudent. A variant considered likely benign has sufficient evidence that a health-care provider can conclude that it is not the cause of the patient’s disorder when combined with other information, for example, if the variant does not segregate in an affected family member and complex inheritance patterns are unlikely. A variant considered benign has sufficient evidence that a health-care provider can conclude that it is not the cause of the patient’s disorder.
医务工作者如何使用基因检测提供的证据来进行医疗管理决策是一个关键问题。目前变异分析是不完善的,报道的变异分类也并不是100%确定的。一般来说,根据推荐的分类方法划分为致病性的变异符合经验数据形成的标准,所以医务工作者可以在临床决策时采用分子检测信息。应尽力避免使用此类信息作为孟德尔疾病的唯一证据,在可能的情况下应与其他临床资料相结合。通常情况下,一个有足够的证据被划分为可能致病的变异,当与可疑疾病的其他证据相结合时,医务工作者可以使用分子检测信息进行临床决策的制定。例如,产前超声可能显示关键的证据,对于产后的病例,其他数据,如酶检测、体格检查,或影像学研究可能最终支持临床决策。然而,推荐进行所有如上所述的可能的后续检测,追踪可能致病变异相关的附加证据的产生,因为这有可能将可能的致病性变异重新归类为致病变异。意义不明确的变异不宜应用于临床决策。应努力将变异分类为致病性或良性。当变异的重新分类正在进行中时,对可疑致病的患者进行额外的监测应审慎。一个有足够证据被考虑为可能良性的变异,医务工作者可以结合其他信息,推断此变异不是该患者致病的原因, 例如,变异并不与家族中的某位患病成员共分离,而且也不太可能是复杂遗传模式。一个有足够证据被考虑为良性的变异,医务工作者可以得出此变异不是该患者致病原因的结论。
How the genetic testing evidence is used is also dependent on the clinical context and indication for testing. In a prenatal diagnostic case where a family is considering irrevocable decisions such as fetal treatment or pregnancy termination, the weight of evidence from the report and other sources such as fetal ultrasound needs to be considered before action is taken. When a genetic test result is the only evidence in a prenatal setting, variants considered likely pathogenic must be explained carefully to families. It is therefore critical for referring healthcare providers to communicate with the clinical laboratory to gain an understanding of how variants are classified to assist in patient counseling and management.
基因检测的证据如何使用也依赖于临床背景和检测指征。在产前诊断的病例中,如果该家庭正在考虑的决定将导致不可逆的后果时,如宫内治疗或终止妊娠等,需要在采取行动之前慎重考虑报告中证据的份量和胎儿超声等其他信息。当基因检测结果是产前检查的唯一证据时,需要向受检家庭慎重解释可能致病的变异。关键是相关的医务工作者应与临床实验室深入沟通,以了解所检测到的变异是如何被分类的,以期为患者提供准确的遗传咨询和临床决策。
8 参考文献(略)
图1
表1 人群数据库,疾病特异性数据库和序列数据库
| 人群数据库 | ||
|---|---|---|
| Exome Aggregation Consortium http://exac.broadinstitute.org/ | 本数据库中的变异信息是通过对61486个独立个体进行全外显子测序获得。同时也是多种特殊疾病和群体遗传学研究中的一部分。库中不包括儿科疾病患者及其相关人群。 | |
| Exome Variant Server http://evs.gs.washington.edu/EVS | 本数据库中的变异信息是通过对几个欧洲和非洲裔大规模人群的全外显子测序获得。当缺乏变异信息时, 默认该数据已覆盖。 | |
| 1000 Genomes Project http://browser.1000genomes.org | 本数据库中的变异信息是通过对26个种群进行低覆盖度的全基因组测序和高覆盖度的靶序列测序获得。本库所提供的信息比Exome Variant Server更具多样性,但也包含有低质量的数据,有些群体中还包含有关联性个体在内。 | |
| dbSNP http://www.ncbi.nlm.nih.gov/snp | 本数据库由多种来源获得的短片段遗传变异(通常≤50 bp)信息组成。库中可能缺乏溯源性研究的细节,也可能包含致病性突变在内。 | |
| dbVar http://www.ncbi.nlm.nih.gov/dbvar | 本数据库由多种来源获得的基因结构变异(通常>50 bp)信息组成。 | |
| 疾病数据库 | ||
| ClinVar http://www.ncbi.nlm.nih.gov/clinvar | 对变异与表型和临床表型之间的关联进行确定的数据库。 | |
| OMIM http://www.omim.org | 本数据库所含人类基因和相关遗传背景,同时具有疾病相关基因遗传变异的代表性样本收录与遗传疾病典型相关的样本变异信息。 | |
| Human Gene Mutation Database http://www.hgmd.org | 本数据库中的变异注释有文献发表。库中大部分内容需付费订阅。 | |
| 其他特殊数据库 | ||
| Human Genome Variation Society http://www.hgvs.org/dblist/dblist.html | 本数据库由人类基因组变异协会(HGVS)开发,提供数千种专门针对人群中的特殊变异进行的注释。数据库很大一部分是基于Leiden Open Variation Database system建立。 | |
| Leiden Open Variation Database http://www.lovd.nl | ||
| DECIPHER http://decipher.sanger.ac.uk | 使用Ensemble基因组浏览器,将基因芯片数据和临床表型进行关联,便于临床医生和研究人员使用的细胞分子遗传学数据库。 | |
| 序列数据库 | ||
| NCBI Genome http://www.ncbi.nlm.nih.gov/genome | 人类全基因组参考序列的来源。 | |
| RefSeqGene http://www.ncbi.nlm.nih.gov/refseq/rsg | 医学相关基因参考序列。 | |
| Locus Reference Genomic (LRG) http://www.lrg-sequence.org | ||
| MitoMap http://www.mitomap.org/MITOMAP/HumanMitoSeq | 对“剑桥版-人类线粒体DNA参考序列”进行修订后形成。 | |
表2 生物信息分析工具
表3 致病变异分级标准
| 致病性证据 | 分类 |
|---|---|
| 非常强 | PVS1:当一个疾病的致病机制为功能丧失(LOF)时,无功能变异(无义突变、移码突变、经典±1或2的剪接突变、起始密码子变异、单个或多个外显子缺失)注:1. 该基因的LOF是否是导致该疾病的明确致病机制(如GFAP、MYH7)2. 3'端末端的功能缺失变异需谨慎解读3.需注意外显子选择性缺失是否影响到蛋白质的完整性4.考虑一个基因存在多种转录本的情况。 |
| 强 | PS1:与先前已确定为致病性的变异有相同的氨基酸改变。例如:同一密码子,G>C或G>T改变均可导致缬氨酸→亮氨酸的改变。注意剪切影响的改变。 |
| PS2:患者的新发变异,且无家族史(经双亲验证)。 注:仅仅确认父母还不够,还需注意捐卵、代孕、胚胎移植的差错等情况。 | |
| PS3:体内、体外功能实验已明确会导致基因功能受损的变异。 注:功能实验需要验证是有效的,且具有重复性与稳定性。 | |
| PS4:变异出现在患病群体中的频率显著高于对照群体。注 1:可选择使用相对风险值或者OR值来评估,建议位点OR大于5.0且置信区间不包括1.0的可列入此项。(详细见指南正文)。2:极罕见的变异在病例对照研究可能无统计学意义,原先在多个具有相同表型的患者中观察到该变异且在对照中未观察到可作为中等水平证据。 | |
| 中等 | PM1:位于热点突变区域,和/或位于已知无良性变异的关键功能域(如酶的活性位点)。 |
| PM2:ESP数据库、千人数据库、EXAC数据库中正常对照人群中未发现的变异(或隐性遗传病中极低频位点)(表6) 注: 高通量测序得到的插入/缺失人群数据质量较差 | |
| PM3:在隐性遗传病中,在反式位置上检测到致病变异。 注:这种情况必须通过患者父母或后代验证。 | |
| PM4:非重复区框内插入/缺失或终止密码子丧失导致的蛋白质长度变化。 | |
| PM5:新的错义突变导致氨基酸变化,此变异之前未曾报道,但是在同一位点,导致另外一种氨基酸的变异已经确认是致病性的,如:现在观察到的是Arg156Cys,而Arg156His是已知致病的。注意剪切影响的改变。 | |
| PM6: 未经父母样本验证的新发变异。 | |
| 支持证据 | PP1:突变与疾病在家系中共分离(在家系多个患者中检测到此变异) 注:如果有更多的证据,可作为更强的证据。 |
| PP2: 对某个基因来说,如果这个基因的错义变异是造成某种疾病的原因,并且这个基因中良性变异所占的比例很小,在这样的基因中所发现的新的错义变异。 | |
| PP3:多种统计方法预测出该变异会对基因或基因产物造成有害的影响,包括保守性预测、进化预测、剪接位点影响等。注:由于做预测时许多生物信息算法使用相同或非常相似的输入,每个算法不应该算作一个独立的标准。PP3在一个任何变异的评估中只能使用一次。 | |
| PP4:变异携带者的表型或家族史高度符合某种单基因遗传疾病。 | |
| PP5:有可靠信誉来源的报告认为该变异为致病的,但证据尚不足以支持进行实验室独立评估。 |
表4 良性变异分类标准
| 良性影响的证据 | 分类 |
|---|---|
| 独立证据 | BA1:ESP数据库、千人数据库、ExAC数据库中等位基因频率>5%的变异。 |
| 强 | BS1:等位基因频率大于疾病发病率。 |
| BS2:对于早期完全外显的疾病,在健康成年人中发现该变异(隐性遗传病发现纯合、显性遗传病发现杂合,或者X连锁半合子)。 | |
| BS3: 在体内外实验中确认对蛋白质功能和剪接没有影响的变异。 | |
| BS4:在一个家系成员中缺乏共分离。 | |
| 注:这部分需要考虑复杂疾病和外显率问题。 | |
| 支持证据 | BP1:已知一个疾病的致病原因是由于某基因的截短变异,在此基因中所发现的错义变异。 |
| BP2:在显性遗传病中又发现了另一条染色体上同一基因的一个已知致病变异,或者是任意遗传模式遗传病中又发现了同一条染色体上同一基因的一个已知致病变异。 | |
| BP3:功能未知重复区域内的缺失/插入,同时没有导致基因编码框改变。 | |
| BP4:多种统计方法预测出该变异会对基因或基因产物无影响,包括保守性预测、进化预测、剪接位点影响等。注:由于做预测时许多生物信息算法使用相同或非常相似的输入,每个算法不应该算作一个独立的标准。BP4在一个任何变异的评估中只能使用一次。 | |
| BP5:在已经有另一分子致病原因的病例中发现的变异。 | |
| BP6:有可靠信誉来源的报告认为该变异为良性的,但证据尚不足以支持进行实验室独立评估。 | |
| BP7:同义变异且预测不影响剪接。 |
表5 遗传变异分类联合标准规则
| 致病的 | (i) 1个非常强(PVS1)和 |
|---|---|
| (a) ≥1个强(PS1-PS4)或 | |
| (b) ≥2个中等(PM1-PM6)或 | |
| (c) 1个中等(PM1-PM6)和1个支持(PP1-PP5)或 | |
| (d) ≥2个支持(PP1-PP5) | |
| (ii) ≥2 个强(PS1-PS4)或 | |
| (iii) 1个强(PS1)和 | |
| (a) ≥3个中等(PM1-PM6)或 | |
| (b) 2个中等(PM1-PM6)和≥2个支持(PP1-PP5)或 | |
| (c) 1个中等(PM1-PM6)和≥4个支持(PP1-PP5) | |
| 可能致病的 | (i) 1个非常强(PVS1)和1个中等(PM1-PM6)或 |
| (ii) 1个强(PS1-PS4)和1-2个中等(PM1-PM6)或 | |
| (iii) 1个强(PS1-PS4)和≥2个支持(PP1-PP5)或 | |
| (iv) ≥3个中等(PM1-PM6)或 | |
| (v) 2个中等(PM1-PM6)和≥2个支持(PP1-PP5)或 | |
| (vi) 1个中等(PM1-PM6)和≥4个支持(PP1-PP5) | |
| 良性的 | (i) 1个独立(BA1)或 |
| (ii) ≥2个强(BS1-BS4) | |
| 可能良性的 | (i) 1个强(BS1-BS4)和1个支持(BP1-BP7)或 |
| (ii) ≥2个支持(BP1-BP7) | |
| 意义不明确的 | (i) 不满足上述标准或 |
| (ii) 良性和致病标准相互矛盾 |